
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 15,075,607 – 15,075,685 |
| Length | 78 |
| Max. P | 0.767361 |

| Location | 15,075,607 – 15,075,685 |
|---|---|
| Length | 78 |
| Sequences | 11 |
| Columns | 87 |
| Reading direction | reverse |
| Mean pairwise identity | 67.12 |
| Shannon entropy | 0.65891 |
| G+C content | 0.45637 |
| Mean single sequence MFE | -14.53 |
| Consensus MFE | -6.86 |
| Energy contribution | -6.52 |
| Covariance contribution | -0.35 |
| Combinations/Pair | 1.33 |
| Mean z-score | -0.96 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.767361 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
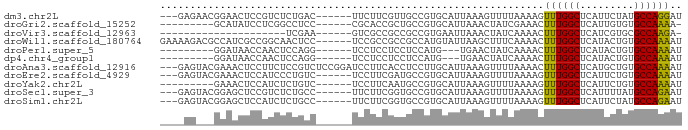
>dm3.chr2L 15075607 78 - 23011544 ---GAGAACGGAACUCCGUCUCUGAC------UUCUUCGUUGCCGUGCAUUAAAGUUUUAAAAGUUUGGCUCAUUCUAUGCCAGGAU ---((((.(((....))))))).(((------((......(((...)))...))))).......((((((.........)))))).. ( -16.20, z-score = -0.17, R) >droGri2.scaffold_15252 905705 71 - 17193109 ---------GCAUAUCCUCGGCCUCC------CGCACCGCUGCCGUGCAUUAAACUAUCGAAACUUUGGCUCAUUGUGUGCCAAAA- ---------((((((....((((...------.((((.......))))......(....).......))))....)))))).....- ( -13.90, z-score = -0.23, R) >droVir3.scaffold_12963 15663197 59 + 20206255 ---------------------UCGAA------GUCGCCGCCGCCGUGAAUUAAACUAUCAAAACUUUGGCUCAUCGUGCGCCAAGA- ---------------------.....------.((((.......))))................((((((.(.....).)))))).- ( -8.70, z-score = 1.26, R) >droWil1.scaffold_180764 3659173 81 - 3949147 GAAAAGACGCCAUCGCCGGCAACUCC------UCCGCCGCCGCCAUGUAUUAAGCUUUCAAAACUUUGGCUCAUACUGUGCCAAAAU ((((......(((.((((((......------...))))..)).)))........)))).....((((((.(.....).)))))).. ( -15.74, z-score = -0.35, R) >droPer1.super_5 2657467 69 - 6813705 ---------GGAUAACCAACUCCAGG------UCCUCCUCCUCCAUG---UGAACUAUCAAAACUUUGGCUCAUACUGUGCCAAAAU ---------(((..(((.......))------)..))).........---..............((((((.(.....).)))))).. ( -13.50, z-score = -2.02, R) >dp4.chr4_group1 2735746 69 + 5278887 ---------GGAUAACCAACUCCAGG------UCCUCCUCCUCCAUG---UGAACUAUCAAAACUUUGGCUCAUACUGUGCCAAAAU ---------(((..(((.......))------)..))).........---..............((((((.(.....).)))))).. ( -13.50, z-score = -2.02, R) >droAna3.scaffold_12916 5369539 84 + 16180835 ---GAGUACGAAACUCCUUCUCCGUCUCCGGAUCCUUCACCUCCUUGCAUUAAAGUUUUAAAACUUUGGCUCAUGCUGUGCCAAAAU ---(.(((((..........((((....))))..............(((((((((((....)))))))....))))))))))..... ( -13.70, z-score = -1.17, R) >droEre2.scaffold_4929 6815489 78 - 26641161 ---GAGUACGAAACUCCAUCCCUGUC------UCCUUCGAUGCCGUGCAUUAAAGUUUUAAAAGUUUGGCUCAUUCUGUGCCAAAAU ---((((.....))))..........------..(((..((((...))))..))).........((((((.(.....).)))))).. ( -15.30, z-score = -2.01, R) >droYak2.chr2L 2052098 72 + 22324452 ---------GAAACUCCAUCUCUGUC------UCCUUCAAUGCCGUGCAUUAAAGUUUUAAAAGUUUGGCUCAUUCUGUGCCAAAAU ---------.................------..(((.(((((...))))).))).........((((((.(.....).)))))).. ( -11.60, z-score = -1.90, R) >droSec1.super_3 1338068 78 - 7220098 ---GAGUACGGAGCUCCGUCUCUGCC------UUCUUCGGUGCCGUGCAUUAAAGUUUUAAAAGUUUGGCUCAUUUUAUGCCAGAAU ---(((.((((....)))))))....------..(((..((((...))))..))).........((((((.........)))))).. ( -20.90, z-score = -1.65, R) >droSim1.chr2L 14820901 78 - 22036055 ---GAGUACGGAGCUCCAUCUCUGCC------UUCUUCGGUGCCGUGCAUUAAAGUUUUAAAAGUUUGGCUCAUUCUAUGCCAGAAU ---..((((((((......))).(((------......)))..)))))................((((((.........)))))).. ( -16.80, z-score = -0.28, R) >consensus ____AG___GGAACUCCAUCUCCGCC______UCCUUCGCUGCCGUGCAUUAAAGUUUUAAAACUUUGGCUCAUUCUGUGCCAAAAU ................................................................((((((.........)))))).. ( -6.86 = -6.52 + -0.35)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:42:06 2011