| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,666,126 – 14,666,229 |
| Length | 103 |
| Max. P | 0.983593 |

| Location | 14,666,126 – 14,666,229 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 66.10 |
| Shannon entropy | 0.68537 |
| G+C content | 0.40225 |
| Mean single sequence MFE | -22.41 |
| Consensus MFE | -9.09 |
| Energy contribution | -10.94 |
| Covariance contribution | 1.85 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.983593 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14666126 103 + 23011544 UGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGCUUUUUCUCCACAA--GCCUCUUUCUCUCGCUUUUUAUGACUACCAUAAAAAUG------- .((((((....((((...((((.((((.....)))).))))..))))..(((((.........))--)))....))))))..((((((((.....))))))))..------- ( -25.60, z-score = -3.44, R) >droSim1.chr2L 14414893 102 + 22036055 UGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGCUUUUUCUCCACA---GCCUCUUUCUCUCGCUUUUUAUGACCACCAUAAAAAUG------- .((((((....((((...((((.((((.....)))).))))..))))..((((..........)---)))....))))))..((((((((.....))))))))..------- ( -24.60, z-score = -2.85, R) >droSec1.super_3 939614 102 + 7220098 UGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGCUUUUUCUCCACA---GCCUCUUUCUCUCGCUUUUUAUGACCACCAUAAAAAUG------- .((((((....((((...((((.((((.....)))).))))..))))..((((..........)---)))....))))))..((((((((.....))))))))..------- ( -24.60, z-score = -2.85, R) >droYak2.chr2L 14772150 88 + 22324452 CGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGC-----------------CUCUUUCUCUCGCUUUUUAUGACUACCAUAAAAGUG------- ((((((((((((.((((..((......))..))))))))).(((.....)))-----------------.....))))))).((((((((.....))))))))..------- ( -20.90, z-score = -2.19, R) >droEre2.scaffold_4929 6413363 102 + 26641161 CGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGCUUUUCUUCCACA---GCCUCUUUCUCUCGCUUUUUAUGACUGCCAUAAAAAUG------- (((((((....((((...((((.((((.....)))).))))..))))..((((..........)---)))....))))))).((((((((.....))))))))..------- ( -25.70, z-score = -2.82, R) >droAna3.scaffold_12916 15011845 103 - 16180835 CAGGAGAUAAAUGUGCAAAUGAUUUGCCAACCGCGAUUUAUGCCACAACGGUUCUUUUUUUA-----GCCUCUUUCUCUUACUCUUUAUGAGCCCCAUAAAGCGAGUG---- .((((((....((((...((((.((((.....)))).))))..))))..((((........)-----)))....))))))((((((((((.....))))))..)))).---- ( -22.80, z-score = -1.55, R) >droWil1.scaffold_180764 912493 92 - 3949147 ------------GUGCAAAUAAUUUGCCAACCGAGAUUUAUCGUUCAACGGUUGUUUCGUCUUUACGGAUUUUUGUUUUUGGCGUUUACGUCUUUGACAAAUUG-------- ------------..((((.....))))((((((.((........))..)))))).(((((....)))))..((((((...((((....))))...))))))...-------- ( -20.30, z-score = -1.27, R) >droGri2.scaffold_15126 991382 98 - 8399593 --------------AGAUAAAUUUAGCCAAUUGCGAUUUCUGCUCAACGUGCUUGCUUUAUAUUCCAGUUUACUUAAGUCUUAGUUGGUGCCAUUAAUGCGCGAGAUUUAGA --------------...(((((((.((((((((.(((((..(...(((...................)))..)..))))).))))))))((((....)).)).))))))).. ( -14.81, z-score = 1.51, R) >consensus CGGGAGAUAAAUGUGCAAAUGAUUUGCCAACUGCGAUUUAUGCCACAACGGCUUUUUCUCCACA___GCCUCUUUCUCUCGCUUUUUAUGACUACCAUAAAAAUG_______ .((((((...........((((.((((.....)))).))))(((.....)))......................))))))...(((((((.....))))))).......... ( -9.09 = -10.94 + 1.85)

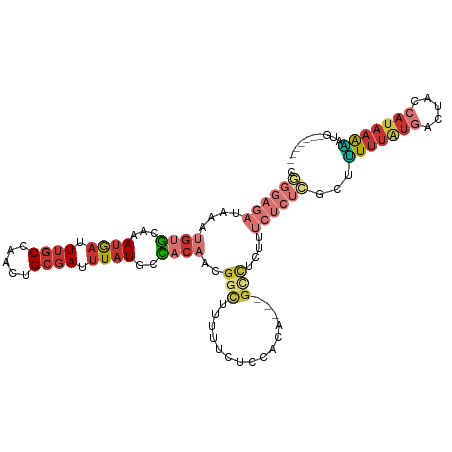
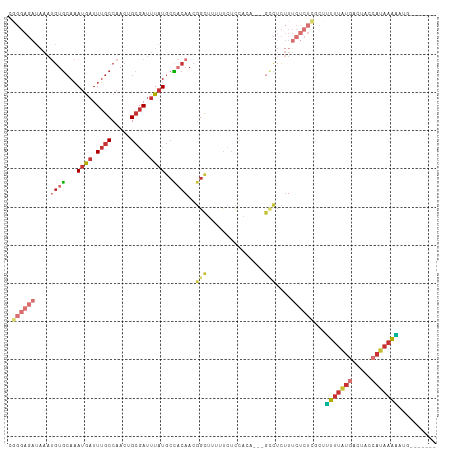
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:41:18 2011