
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,586,745 – 14,586,796 |
| Length | 51 |
| Max. P | 0.731252 |

| Location | 14,586,745 – 14,586,796 |
|---|---|
| Length | 51 |
| Sequences | 9 |
| Columns | 56 |
| Reading direction | forward |
| Mean pairwise identity | 86.56 |
| Shannon entropy | 0.26635 |
| G+C content | 0.50576 |
| Mean single sequence MFE | -13.06 |
| Consensus MFE | -9.79 |
| Energy contribution | -9.69 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.615195 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14586745 51 + 23011544 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCU-----UCUGAGACGCCCACUAU (((.(((.....)))))).....((.(((((((.-----...)))).))))).... ( -12.70, z-score = -2.04, R) >droSim1.chr2L 14337368 51 + 22036055 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCU-----UCUGAGACGCCCACUAU (((.(((.....)))))).....((.(((((((.-----...)))).))))).... ( -12.70, z-score = -2.04, R) >droSec1.super_3 859983 51 + 7220098 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCU-----UCUGAGACGCCCACUAU (((.(((.....)))))).....((.(((((((.-----...)))).))))).... ( -12.70, z-score = -2.04, R) >droYak2.chr2L 14698472 53 + 22324452 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCUACU--UCUGAGACGCCCACAA- (((.(((.....))))))....(((((((((((....--...)))).).))))))- ( -13.80, z-score = -2.32, R) >droEre2.scaffold_4929 6341131 51 + 26641161 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCU-----UCUGAGACGCCCACUAU (((.(((.....)))))).....((.(((((((.-----...)))).))))).... ( -12.70, z-score = -2.04, R) >droAna3.scaffold_12916 14947906 52 - 16180835 GUCCUCGUUUGCUGGGACUAAUUUGUGGCCCUCGA----CUAAAGACGCCCACUAA (((((.(....).)))))......((((.(.((..----.....)).).))))... ( -12.70, z-score = -1.29, R) >dp4.chr4_group1 2022971 56 - 5278887 AUCCUCGUUUGCUGGGACUAAUUUGUGGCUCUCCUCUCGACUGAGACGCCCACAGU .((((.(....).)))).....(((((((((((.........)))).).)))))). ( -13.40, z-score = -0.43, R) >droPer1.super_5 1966660 56 + 6813705 AUCCUCGUUUGCUGGGACUAAUUUGUGGCUCUCCUCUCGACUGAGACGCCCACAGU .((((.(....).)))).....(((((((((((.........)))).).)))))). ( -13.40, z-score = -0.43, R) >droWil1.scaffold_180764 2937008 51 + 3949147 GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCA-----ACUGGGACGCCCACUAA (((((.((....((((((((.....))).)))))-----)).)))))......... ( -13.40, z-score = -1.63, R) >consensus GUCCUCGUUUGCUGAGACUAAUUUGUGGCUCUCU_____UCUGAGACGCCCACUAU ...((((.....))))........(((((((((.........)))).).))))... ( -9.79 = -9.69 + -0.10)



| Location | 14,586,745 – 14,586,796 |
|---|---|
| Length | 51 |
| Sequences | 9 |
| Columns | 56 |
| Reading direction | reverse |
| Mean pairwise identity | 86.56 |
| Shannon entropy | 0.26635 |
| G+C content | 0.50576 |
| Mean single sequence MFE | -12.81 |
| Consensus MFE | -11.01 |
| Energy contribution | -10.53 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.86 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.731252 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
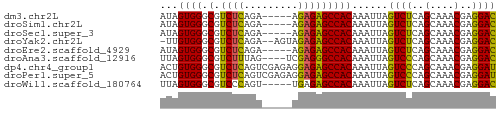
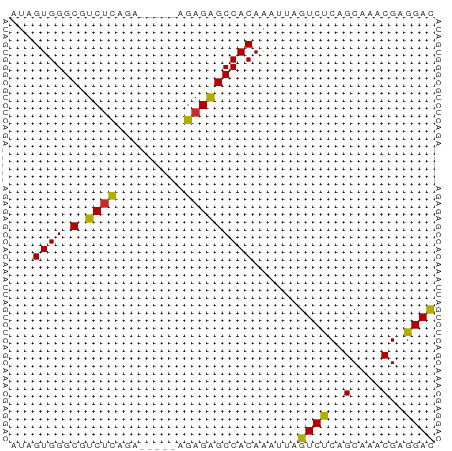
>dm3.chr2L 14586745 51 - 23011544 AUAGUGGGCGUCUCAGA-----AGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ...((((.(.((((...-----.))))))))).......((((.......)))).. ( -12.30, z-score = -1.69, R) >droSim1.chr2L 14337368 51 - 22036055 AUAGUGGGCGUCUCAGA-----AGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ...((((.(.((((...-----.))))))))).......((((.......)))).. ( -12.30, z-score = -1.69, R) >droSec1.super_3 859983 51 - 7220098 AUAGUGGGCGUCUCAGA-----AGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ...((((.(.((((...-----.))))))))).......((((.......)))).. ( -12.30, z-score = -1.69, R) >droYak2.chr2L 14698472 53 - 22324452 -UUGUGGGCGUCUCAGA--AGUAGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC -((((((.(.((((...--....))))))))))).....((((.......)))).. ( -13.40, z-score = -1.62, R) >droEre2.scaffold_4929 6341131 51 - 26641161 AUAGUGGGCGUCUCAGA-----AGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ...((((.(.((((...-----.))))))))).......((((.......)))).. ( -12.30, z-score = -1.69, R) >droAna3.scaffold_12916 14947906 52 + 16180835 UUAGUGGGCGUCUUUAG----UCGAGGGCCACAAAUUAGUCCCAGCAAACGAGGAC ...((((.(.((.....----..)).).))))......((((..(....)..)))) ( -13.90, z-score = -1.43, R) >dp4.chr4_group1 2022971 56 + 5278887 ACUGUGGGCGUCUCAGUCGAGAGGAGAGCCACAAAUUAGUCCCAGCAAACGAGGAU ..(((((.(.((((....)))).)....))))).....((((..(....)..)))) ( -13.90, z-score = -0.48, R) >droPer1.super_5 1966660 56 - 6813705 ACUGUGGGCGUCUCAGUCGAGAGGAGAGCCACAAAUUAGUCCCAGCAAACGAGGAU ..(((((.(.((((....)))).)....))))).....((((..(....)..)))) ( -13.90, z-score = -0.48, R) >droWil1.scaffold_180764 2937008 51 - 3949147 UUAGUGGGCGUCCCAGU-----UGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ....((((...))))((-----(((((............))))))).......... ( -11.00, z-score = -0.73, R) >consensus AUAGUGGGCGUCUCAGA_____AGAGAGCCACAAAUUAGUCUCAGCAAACGAGGAC ...((((.(.((((.........)))))))))......((((..(....)..)))) (-11.01 = -10.53 + -0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:55 2011