
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,559,361 – 14,559,417 |
| Length | 56 |
| Max. P | 0.723590 |

| Location | 14,559,361 – 14,559,417 |
|---|---|
| Length | 56 |
| Sequences | 11 |
| Columns | 58 |
| Reading direction | reverse |
| Mean pairwise identity | 88.07 |
| Shannon entropy | 0.28116 |
| G+C content | 0.47771 |
| Mean single sequence MFE | -15.57 |
| Consensus MFE | -9.80 |
| Energy contribution | -10.62 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.723590 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14559361 56 - 23011544 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >droSim1.chr2L 14309796 56 - 22036055 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >droSec1.super_3 822711 56 - 7220098 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >droYak2.chr2L 14670451 56 - 22324452 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >droEre2.scaffold_4929 6314459 56 - 26641161 UGGGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG (((((((((((((.--(.....).)))).)))))).)))................... ( -17.80, z-score = -2.14, R) >droAna3.scaffold_12916 14922546 56 + 16180835 UGCGACUGCCAUCG--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG ...(.(((((((((--(.....)))))).)))).)....................... ( -14.10, z-score = -0.90, R) >droPer1.super_5 1932804 56 - 6813705 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >dp4.chr4_group1 1989027 56 + 5278887 UGUGGCUGCCAUCU--CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG .((((((((((((.--(.....).)))).))))))))..................... ( -18.90, z-score = -3.15, R) >droWil1.scaffold_180764 1053574 53 + 3949147 -----CUGCCUUUUGCCGCUGACUGAUGUGAGGCCAUCAAUCUAAACUCAUUAUUUUG -----..((((.....(((........)))))))........................ ( -5.40, z-score = 0.59, R) >droMoj3.scaffold_6500 16522556 56 + 32352404 UUUCGUCUCCACAU--CGUUGACUGAUGUGUGGCCAUCAAUCUAAACUCUUCAUUAUG ....(((..(((((--((.....))))))).)))........................ ( -9.40, z-score = -1.50, R) >droGri2.scaffold_15126 925561 58 - 8399593 UGUGGCAGCUCCGCCUCGCUGACCGAUGUGUGGCCAUCAAUCUAAACUCUUCAUUUUG ...((((((........)))).))((((......)))).................... ( -11.20, z-score = -0.20, R) >consensus UGUGGCUGCCAUCU__CGCUGAGUGAUGUGCGGCCAUCAAUCUAAACUCAUCAUUUUG ...((((((((((...........)))).))))))....................... ( -9.80 = -10.62 + 0.81)

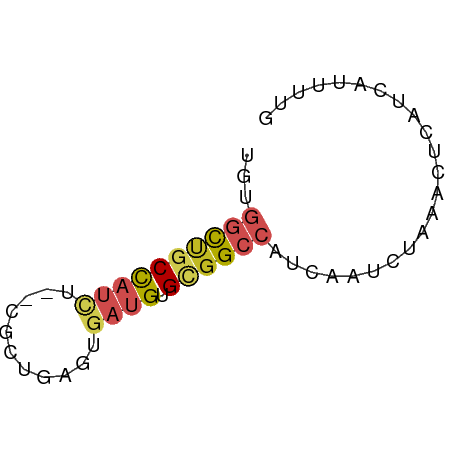

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:51 2011