| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,552,971 – 14,553,064 |
| Length | 93 |
| Max. P | 0.900591 |

| Location | 14,552,971 – 14,553,064 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 63.65 |
| Shannon entropy | 0.68543 |
| G+C content | 0.49873 |
| Mean single sequence MFE | -20.82 |
| Consensus MFE | -10.44 |
| Energy contribution | -9.59 |
| Covariance contribution | -0.85 |
| Combinations/Pair | 1.55 |
| Mean z-score | -0.97 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.900591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
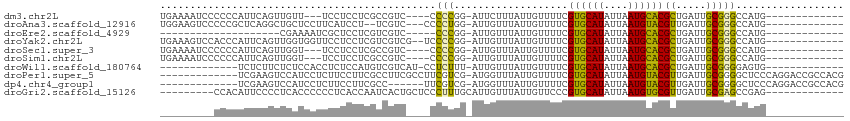
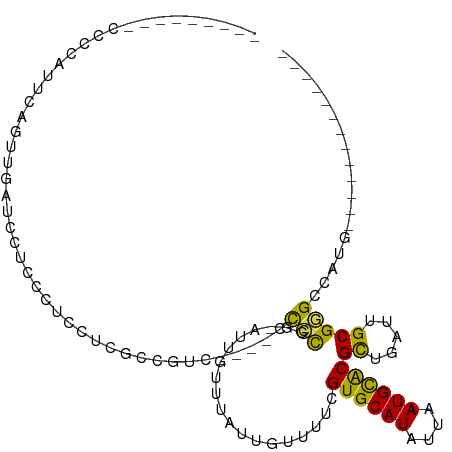
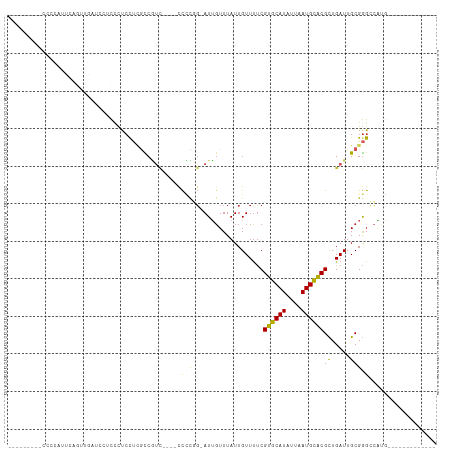
>dm3.chr2L 14552971 93 + 23011544 UGAAAAUCCCCCCAUUCAGUUGUU---UCCUCCUCGCCGUC----CCCCGG-AUUCUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG------------- .........(((((.(((((....---.........(((..----...)))-................((((((....))))))))))).)).))).....------------- ( -22.30, z-score = -2.29, R) >droAna3.scaffold_12916 14915947 95 - 16180835 UGGAAGUCCCCCGCUCAGGCUGCUCCUUCAUCCU--UCGUC---CCCCUGG-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGUACGUUGAUUGCGGGCCAUG------------- .((....))((((((((((..((...........--..)).---..)))))-..........(((..(((((((....)))))))..))).))))).....------------- ( -22.22, z-score = -0.52, R) >droEre2.scaffold_4929 6308361 75 + 26641161 --------------------CGAAAAUCGCUCCUCGUCGUC-----CCCGG-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG------------- --------------------........((((...((((((-----....)-)).............(((((((....)))))))..)))...))))....------------- ( -16.00, z-score = 0.36, R) >droYak2.chr2L 14663848 98 + 22324452 UGAAAGUCCACCCAUUCAGUUGGUGGUUCCUCCUCGUCGUCG--UCCCCGG-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG------------- .((....(((((.........)))))....))(((((.((((--((....)-)).............(((((((....)))))))..))).))))).....------------- ( -24.30, z-score = -0.44, R) >droSec1.super_3 816373 93 + 7220098 UGAAAAUCCCCCCAUUCAGUUGGU---UCCUCCUCGCCGUC----CCCCGG-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG------------- .........(((((.(((((.(((---........)))(((----(...))-))..............((((((....))))))))))).)).))).....------------- ( -22.80, z-score = -1.35, R) >droSim1.chr2L 14303457 93 + 22036055 UGAAAAUCCCCCCAUUCAGUUGGU---UCCUCCUCGCCGUC----CCCCGG-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG------------- .........(((((.(((((.(((---........)))(((----(...))-))..............((((((....))))))))))).)).))).....------------- ( -22.80, z-score = -1.35, R) >droWil1.scaffold_180764 1067385 86 - 3949147 -------------UCUCUUCUCUCCACCUCUCCAUGUCGUCAU-CCUCUUU-AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGGAGUG------------- -------------..............((((((..........-.......-................((((((....))))))((.....))))))))..------------- ( -19.30, z-score = -2.80, R) >droPer1.super_5 1926426 100 + 6813705 -------------UCGAAGUCCAUCCUCUUCCUUCGCCUUCGCCUUCGUCG-AUGGUUUAUUGUUUUCGUGCAUAUUAAUGUACGUUGAUUGCGGGGCUCCCAGGACCGCCACG -------------.....((((............(((..(((((.......-..)))..........(((((((....)))))))..))..)))((....)).))))....... ( -22.40, z-score = 0.33, R) >dp4.chr4_group1 1982625 94 - 5278887 -------------UCGAAGUCCAUCCUCUUCCUUCGCC------UUCGUCG-AUGGUUUAUUGUUUUCGUGCAUAUUAAUGUACGUUGAUUGCGGGGCUCCCAGGACCGCCACG -------------.(((((............)))))..------..(((.(-.((((((...(((..(((((((....)))))))..)))...(((...))).))))))).))) ( -20.80, z-score = 0.38, R) >droGri2.scaffold_15126 915846 92 + 8399593 ---------CCACAUUCCCCUCACCCCCCUCACCAAUCACUGCUCCCUUUGCAUUGUUUAUUGUUCCCGUGCAUAUUAAUGUGCGUUGAUUGCGAGCCGAG------------- ---------...................(((..((((((.(((.......)))..............((..(((....)))..)).)))))).))).....------------- ( -15.30, z-score = -2.00, R) >consensus _________CCCCAUUCAGUUGAUCCUCCCUCCUCGCCGUC____CCCCGG_AUUGUUUAUUGUUUUCGUGCAUAUUAAUGCACGCUGAUUGCGGGCCAUG_____________ ..............................................(((...................((((((....))))))((.....))))).................. (-10.44 = -9.59 + -0.85)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:49 2011