
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,307,868 – 1,307,958 |
| Length | 90 |
| Max. P | 0.515958 |

| Location | 1,307,868 – 1,307,958 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 71.84 |
| Shannon entropy | 0.56078 |
| G+C content | 0.51426 |
| Mean single sequence MFE | -29.18 |
| Consensus MFE | -11.01 |
| Energy contribution | -13.17 |
| Covariance contribution | 2.16 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.515958 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 1307868 90 + 23011544 ACUAGCAUUAAGGUAAAUCCUUU-CAAGCACUUCAAGUGU-------UUGUCGGCGGUGUCGAGGGUGGGCGUGUUUGACAAACAC-CCCAUCCACGCA-- ....((...((((.....)))).-(((((((.....))))-------)))(((((...)))))(((((((.(((((.....)))))-)))))))..)).-- ( -33.10, z-score = -3.11, R) >droAna3.scaffold_12916 4771287 91 + 16180835 ACUAGCAUUAAGUUAAAUCCUUUUCAAGCACUUCAAGUGU-------UUGCCGGGGGUGUUGACGGUGGGCGUGUCCGACAAACAC-CCCAUAGACGCA-- ....((.....(((((((((((..(((((((.....))))-------)))..)))))).))))).(((((.((((.......))))-)))))....)).-- ( -31.50, z-score = -2.28, R) >droYak2.chr2L 1283772 90 + 22324452 AGCAGCAUUAAGGUAAAUCCUUU-CAAGCACUUCAAGUGU-------UUGUCGGCGGCGUUGAAGGUGGGCAUGUUUGACAAACAC-CCCAUCGACGCA-- ....((...((((.....)))).-(((((((.....))))-------)))...)).(((((((...((((..((((.....)))).-))))))))))).-- ( -27.80, z-score = -1.45, R) >droEre2.scaffold_4929 1346557 90 + 26641161 ACUAGCAUUAAGGUAAAUCCCUU-CAAGCACUUCAAGUGU-------UUGUCGGCGGCGUGGAAGGUGGGCGUGUUUGGCAAACAC-CCCAUCGACGCA-- ....((...((((......))))-(((((((.....))))-------)))...)).((((....((((((.(((((.....)))))-)))))).)))).-- ( -31.30, z-score = -1.69, R) >droSec1.super_14 1257502 90 + 2068291 ACUAGCAUUAAGGCAAAUCCUUU-CAAGCACUUCAAGUGU-------UUGUCGGCGGUGUUGAGGGUGGGCGUGUUUGACAAACAC-CCCAUCGACGCA-- ....((.((((.((....((...-(((((((.....))))-------)))..))..)).)))).((((((.(((((.....)))))-))))))...)).-- ( -30.60, z-score = -1.86, R) >droSim1.chr2L 1279328 90 + 22036055 ACUAGCAUUAAGGUAAAUCCUUU-CAAGCACUUCAAGUGU-------UUGUCGGCGGCGUUGAGGGUGGGCGUGUUUGACAAACAC-CCCAUCGACGCA-- ....((...((((.....)))).-(((((((.....))))-------)))...)).(((((((...((((.(((((.....)))))-))))))))))).-- ( -32.50, z-score = -2.67, R) >droPer1.super_8 997109 88 + 3966273 -------------UAAAUCCUUUUCGAACACUUGGAUUGUGUGUGUCUUGUCGAGGGUGUUGACGGUGGGCGUAUUUGACAAACACACCCCCUGGGGCCCG -------------..(((((.............)))))..((((((.((((((((..((((.......))))..))))))))))))))((....))..... ( -23.12, z-score = 0.98, R) >droMoj3.scaffold_6500 28721816 70 - 32352404 AUCAGCAUACAGCUAACCCCAUG-CUCGCACCAGGAGGC--------------GGGCUGUUAGAAGUGGGCGUAACCUGUACGCC---------------- ....((.(((((...((((((((-(((((........))--------------))))........))))).))...))))).)).---------------- ( -23.50, z-score = -0.56, R) >consensus ACUAGCAUUAAGGUAAAUCCUUU_CAAGCACUUCAAGUGU_______UUGUCGGCGGUGUUGAAGGUGGGCGUGUUUGACAAACAC_CCCAUCGACGCA__ ............((.............((((.....)))).............)).((((....((((((.(((((.....))))).)))))).))))... (-11.01 = -13.17 + 2.16)


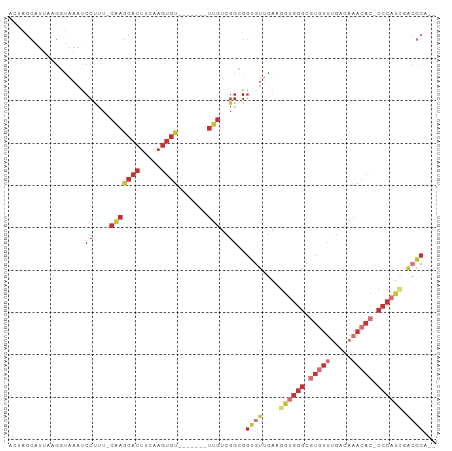
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:18 2011