
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,511,076 – 14,511,169 |
| Length | 93 |
| Max. P | 0.785979 |

| Location | 14,511,076 – 14,511,169 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 87.89 |
| Shannon entropy | 0.23221 |
| G+C content | 0.52865 |
| Mean single sequence MFE | -33.44 |
| Consensus MFE | -29.33 |
| Energy contribution | -29.76 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.785979 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14511076 93 + 23011544 ---------CUGGCG---GUGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAGC ---------.....(---.(((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..))))))).)... ( -35.80, z-score = -1.77, R) >droSim1.chr2L 14258979 93 + 22036055 ---------CUGGCG---GUGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAGC ---------.....(---.(((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..))))))).)... ( -35.80, z-score = -1.77, R) >droSec1.super_3 774227 93 + 7220098 ---------CUGGCG---GUGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAGC ---------.....(---.(((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..))))))).)... ( -35.80, z-score = -1.77, R) >droYak2.chr2L 14620221 83 + 22324452 ---------CUGGCG---GUGCUGUCGUCUCUCACAUGUCAGCGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGA---------- ---------..((((---.(((((.(((.......))).)))).).))))(((((..((.(((((.-....))))).))..)))))..........---------- ( -24.10, z-score = 0.42, R) >droEre2.scaffold_4929 6265146 93 + 26641161 ---------CUGGCG---GUGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAGC ---------.....(---.(((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..))))))).)... ( -35.80, z-score = -1.77, R) >droAna3.scaffold_12916 14874955 93 - 16180835 ---------CUGGCG---GAGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAAC ---------...((.---..((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..)))))).))... ( -34.20, z-score = -1.76, R) >dp4.chr4_group1 1929084 94 - 5278887 ---------UCAUCGU--ACGUUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGCGAAAUGAAGCAACGAGGUGACAGCAGCAAC ---------.......--..((((((..(((......(((....)))((.(((((..((.(((((.-....))))).))..))))).)))))..))))))...... ( -31.80, z-score = -1.56, R) >droPer1.super_5 1873559 95 + 6813705 ---------UCAUCGU--ACGUUGUCGUCUCUCACAUGUCAACAGGCGUCGCUUUGCUUGGCAUUUCUUGUGAUGCGAAAUGAAGCAACGAGGUGACAGCAGCAAC ---------.......--..((((((..(((......(((....)))((.(((((..((.(((((......))))).))..))))).)))))..))))))...... ( -32.10, z-score = -1.68, R) >droVir3.scaffold_12723 3134144 99 - 5802038 CUGGCAGAGCCGGCGACGUCCUCGUCGUCUCUCACAUGUCAACGCGCGUCGCUUUGCUUGGCAUUU-UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGC------ ...((.(((..(((((((....))))))).)))...(((((.(...(((.(((((..((.(((((.-....))))).))..))))).))).).)))))))------ ( -35.60, z-score = -0.85, R) >consensus _________CUGGCG___GUGCUGUCGUCUCUCACAUGUCAACGGGCGUCGCUUUGCUUGGCAUUU_UUGUGAUGUGAAAUGAAGCAACGAGGUGACAGCAGCAGC ....................((((((..(((......(((....)))((.(((((..((.(((((......))))).))..))))).)))))..))))))...... (-29.33 = -29.76 + 0.42)


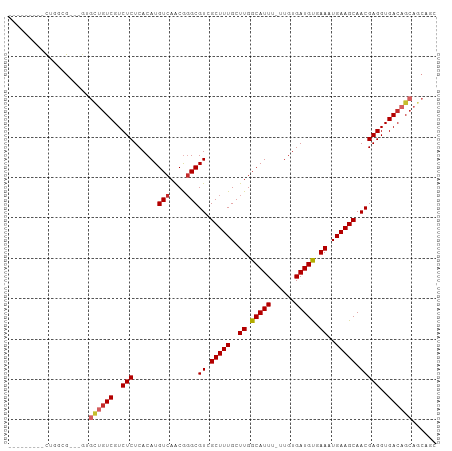
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:32 2011