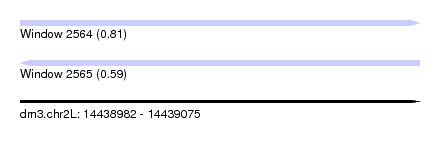

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,438,982 – 14,439,075 |
| Length | 93 |
| Max. P | 0.805003 |

| Location | 14,438,982 – 14,439,075 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 84.86 |
| Shannon entropy | 0.29277 |
| G+C content | 0.48312 |
| Mean single sequence MFE | -27.22 |
| Consensus MFE | -22.51 |
| Energy contribution | -22.07 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.805003 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
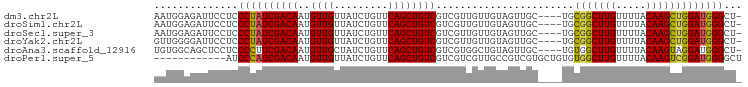
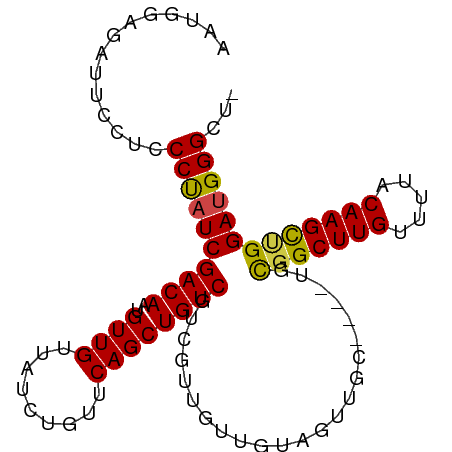
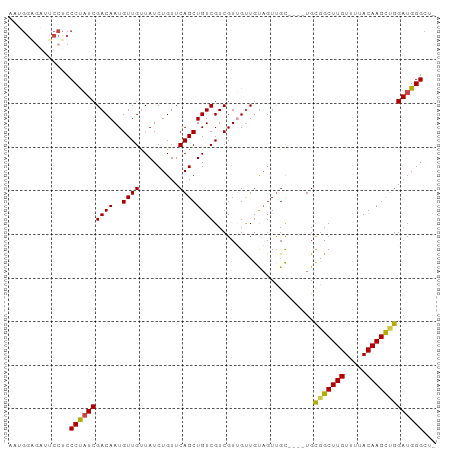
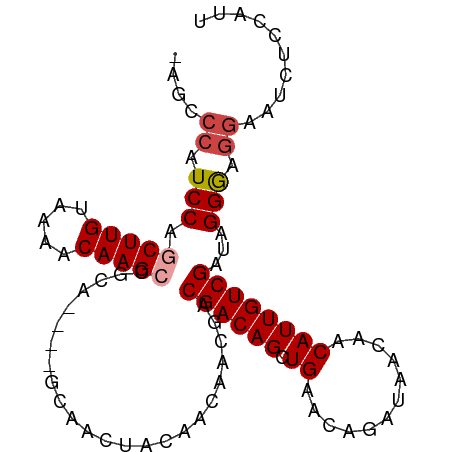
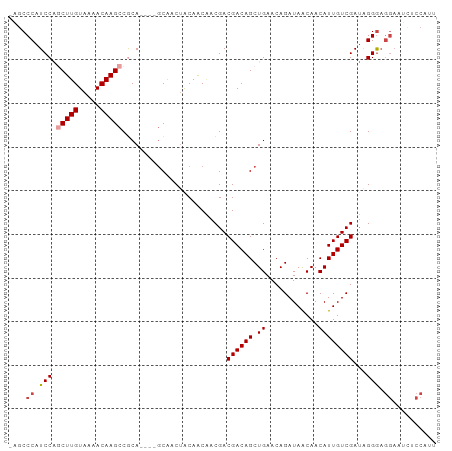
>dm3.chr2L 14438982 93 + 23011544 AAUGGAGAUUCCUCCCUAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUUGUUGUAGUUGC----UGCGGCUUGUUUUACAAGCUGGAUGGGCU- ...((((....))))((((((((((((.((.((.((....)).)).)).)))))))........----..(((((((.....))))))))))))...- ( -26.20, z-score = -1.37, R) >droSim1.chr2L 14187726 93 + 22036055 AAUGGAGAUUCCUCCCUAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUUGUUGUAGUUGC----UGCGGCUUGUUUUACAAGCUGGAUGGGCU- ...((((....))))((((((((((((.((.((.((....)).)).)).)))))))........----..(((((((.....))))))))))))...- ( -26.20, z-score = -1.37, R) >droSec1.super_3 698161 93 + 7220098 AAUGGAGAUUCCUCCCUAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUUGUUGUAGUUGC----UGCGGCUUGUUUUACAAGCUGGAUGGGCU- ...((((....))))((((((((((((.((.((.((....)).)).)).)))))))........----..(((((((.....))))))))))))...- ( -26.20, z-score = -1.37, R) >droYak2.chr2L 14543786 93 + 22324452 GUUGGGGAUUCCUCCCUAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUUGUUGUAGUUGC----UGCGGCUUGUUUUACAAGCUGGAUGGGCU- (.((((((....)))))).)((((....)))).(((((((((((((.......(((((((...)----)))))).......)).)))))))))))..- ( -27.74, z-score = -1.53, R) >droAna3.scaffold_12916 14799672 93 - 16180835 UGUGGCAGCUCCUCCCCUUCGACAAUGUUGCUAUCUGUUCAGCUGUCGUCGUGGCUGUAGUUGC----UGUGGCUUGUUUUACAAGUAGGAUGGGCU- .(..((((((.(..(((..(((((.((..((.....)).))..)))))..).))..).))))))----..)((((..(((((....)))))..))))- ( -30.60, z-score = -1.31, R) >droPer1.super_5 1763520 86 + 6813705 ------------AUCCCAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUCGUUGCCGUCGUGCUGUGUGGCUUGUUUUACAAGUCGGAUGGGGCU ------------.(((((((((((....)))).......((((((.((.((....)).)).)).))))..(((((((.....)))))))))))))).. ( -26.40, z-score = -1.64, R) >consensus AAUGGAGAUUCCUCCCUAUCGACAAUGUUGUUAUCUGUUCAGCUGUCGUCGUUGUUGUAGUUGC____UGCGGCUUGUUUUACAAGCUGGAUGGGCU_ ..............((((((((((..((((.........)))))))).......................(((((((.....)))))))))))))... (-22.51 = -22.07 + -0.44)
| Location | 14,438,982 – 14,439,075 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 84.86 |
| Shannon entropy | 0.29277 |
| G+C content | 0.48312 |
| Mean single sequence MFE | -19.74 |
| Consensus MFE | -13.77 |
| Energy contribution | -14.30 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.592803 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
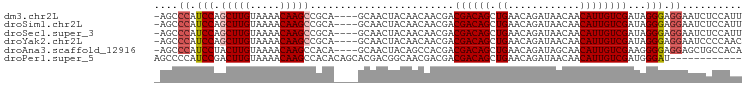
>dm3.chr2L 14438982 93 - 23011544 -AGCCCAUCCAGCUUGUAAAACAAGCCGCA----GCAACUACAACAACGACGACAGCUGAACAGAUAACAACAUUGUCGAUAGGGAGGAAUCUCCAUU -..((((((..(((((.....)))))..((----((...................))))....(((((.....)))))))).))).((.....))... ( -18.61, z-score = -1.41, R) >droSim1.chr2L 14187726 93 - 22036055 -AGCCCAUCCAGCUUGUAAAACAAGCCGCA----GCAACUACAACAACGACGACAGCUGAACAGAUAACAACAUUGUCGAUAGGGAGGAAUCUCCAUU -..((((((..(((((.....)))))..((----((...................))))....(((((.....)))))))).))).((.....))... ( -18.61, z-score = -1.41, R) >droSec1.super_3 698161 93 - 7220098 -AGCCCAUCCAGCUUGUAAAACAAGCCGCA----GCAACUACAACAACGACGACAGCUGAACAGAUAACAACAUUGUCGAUAGGGAGGAAUCUCCAUU -..((((((..(((((.....)))))..((----((...................))))....(((((.....)))))))).))).((.....))... ( -18.61, z-score = -1.41, R) >droYak2.chr2L 14543786 93 - 22324452 -AGCCCAUCCAGCUUGUAAAACAAGCCGCA----GCAACUACAACAACGACGACAGCUGAACAGAUAACAACAUUGUCGAUAGGGAGGAAUCCCCAAC -..((((((..(((((.....)))))..((----((...................))))....(((((.....)))))))).))).((.....))... ( -18.01, z-score = -1.48, R) >droAna3.scaffold_12916 14799672 93 + 16180835 -AGCCCAUCCUACUUGUAAAACAAGCCACA----GCAACUACAGCCACGACGACAGCUGAACAGAUAGCAACAUUGUCGAAGGGGAGGAGCUGCCACA -(((((.(((..((((.....))))((...----((.......))..((((((..((((......))))....))))))..))))))).)))...... ( -22.00, z-score = -0.96, R) >droPer1.super_5 1763520 86 - 6813705 AGCCCCAUCCGACUUGUAAAACAAGCCACACAGCACGACGGCAACGACGACGACAGCUGAACAGAUAACAACAUUGUCGAUGGGAU------------ ...((((((.(.((((.....)))).)...((((.((.((....)).))......))))....(((((.....)))))))))))..------------ ( -22.60, z-score = -3.12, R) >consensus _AGCCCAUCCAGCUUGUAAAACAAGCCGCA____GCAACUACAACAACGACGACAGCUGAACAGAUAACAACAUUGUCGAUAGGGAGGAAUCUCCAUU ....((.(((.(((((.....)))))........................((((((.((............))))))))...))).)).......... (-13.77 = -14.30 + 0.53)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:17 2011