
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,306,099 – 1,306,191 |
| Length | 92 |
| Max. P | 0.617903 |

| Location | 1,306,099 – 1,306,191 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 76.79 |
| Shannon entropy | 0.44919 |
| G+C content | 0.41093 |
| Mean single sequence MFE | -24.12 |
| Consensus MFE | -9.77 |
| Energy contribution | -10.12 |
| Covariance contribution | 0.35 |
| Combinations/Pair | 1.37 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.617903 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 1306099 92 + 23011544 CGCCUAACGGAUUUUCCUC---AAGGG-AACUUUUCC---CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAAAAA---------- ..((....)).........---(((((-(.....)))---)))((((((...))))))........(((((.....)))))..................---------- ( -21.00, z-score = -2.31, R) >droSim1.chr2L 1277610 89 + 22036055 CGCCUAACGGAUUUUCCUC---AAGGG-AACUUUUCC---CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAA------------- ..((....)).........---(((((-(.....)))---)))((((((...))))))........(((((.....)))))...............------------- ( -21.00, z-score = -2.44, R) >droSec1.super_14 1255779 89 + 2068291 CGCCUAACGGAUUUUCCUC---AAGGG-AACUUUUCC---CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAA------------- ..((....)).........---(((((-(.....)))---)))((((((...))))))........(((((.....)))))...............------------- ( -21.00, z-score = -2.44, R) >droEre2.scaffold_4929 1344837 89 + 26641161 CGCCCAACGGAUUUUCCUC---AAGGG-AACUUUUCC---CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCCUUUUGCCAACAACUUGUUAAAAAA------------- ........((.....))..---(((((-(.....)))---)))((((((...))))))........(((((.....)))))...............------------- ( -18.70, z-score = -1.29, R) >droYak2.chr2L 1281963 89 + 22324452 CGCCUAACGGAUUUUCCUC---AAGGG-AACUUUUCC---CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAA------------- ..((....)).........---(((((-(.....)))---)))((((((...))))))........(((((.....)))))...............------------- ( -21.00, z-score = -2.44, R) >droAna3.scaffold_12916 4769638 103 + 16180835 CGGCUAACGGAUUUUCCUC---AAGGGGAAGUUUUCC---CUUUAUGGGCAUGCCAUAUAAAGUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAAAAAAAAAUAAACC ....((((((....(((..---(((((((.....)))---))))..)))....)).....(((((.(((((.....))))).))))))))).................. ( -27.20, z-score = -2.13, R) >dp4.chr4_group3 9793567 93 + 11692001 CGGCUAACGGAUUUUCCUC---AAGGGGAAGUUUUCC---CUUUAUGGGAAUGCCAUAUAAAUUUCUUGGCUUUUGGCCAACAACUUGUUUGGCCAAAA---------- .((((((..(((((((((.---...)))))((.((((---(.....))))).))......))))..))))))(((((((((........))))))))).---------- ( -32.70, z-score = -3.19, R) >droPer1.super_8 994233 93 + 3966273 CGGCUAACGGAUUUUCCUC---AAGGGGAAGUUUUCC---CUUUAUGGGAAUGCCAUAUAAAUUUCUUGGCUUUUGGCCAACAACUUGUUUGGCCAAAA---------- .((((((..(((((((((.---...)))))((.((((---(.....))))).))......))))..))))))(((((((((........))))))))).---------- ( -32.70, z-score = -3.19, R) >droWil1.scaffold_180708 2943248 79 + 12563649 ------ACGGAUUUUUCUUCUUAAGGCCAAGUUUUGC---C-----------ACCAUAUAAAGUGCUUGGCUUUGGGCCAACAACUUGUAUGGAAAAAA---------- ------..................(((........))---)-----------.((((((((..((.(((((.....)))))))..))))))))......---------- ( -21.50, z-score = -1.54, R) >droGri2.scaffold_15252 10385975 90 + 17193109 ------ACGAAUAUCACCC---AGGGGCAAGUUUGCCACACUACGGGGCAUGGUCUUUUAAUGCGGCUGGCUUACGGCCAACAACUUGUACAAGCGGCA---------- ------..........((.---...))......((((.(..((((((((((.........))))(((((.....))))).....))))))...).))))---------- ( -24.40, z-score = 0.32, R) >consensus CGCCUAACGGAUUUUCCUC___AAGGG_AACUUUUCC___CUUUAUGGCCAUGCCAUAUAAACUUCUUGGCUUUUUGCCAACAACUUGUUAAAAAAAAA__________ .....(((((....((((.....))))................(((((.....)))))........(((((.....)))))....)))))................... ( -9.77 = -10.12 + 0.35)
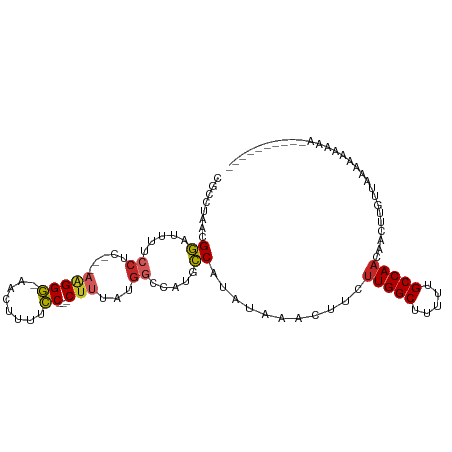


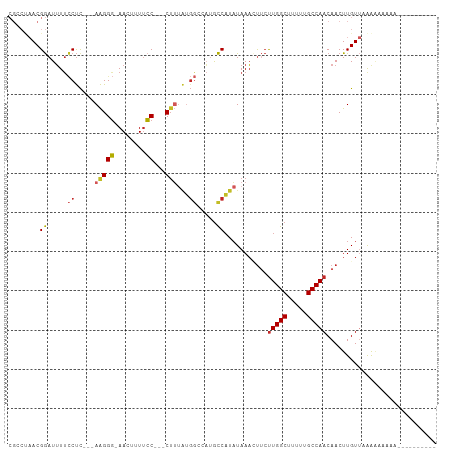
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:16 2011