| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,411,554 – 14,411,654 |
| Length | 100 |
| Max. P | 0.973621 |

| Location | 14,411,554 – 14,411,654 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 68.57 |
| Shannon entropy | 0.65592 |
| G+C content | 0.40932 |
| Mean single sequence MFE | -27.32 |
| Consensus MFE | -10.46 |
| Energy contribution | -11.36 |
| Covariance contribution | 0.90 |
| Combinations/Pair | 1.73 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.973621 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
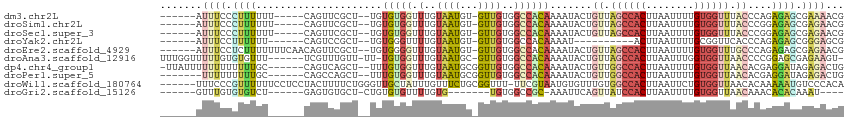
>dm3.chr2L 14411554 100 - 23011544 ------AUUUCCCUUUUUU-----CAGUUCGCU--UGUGUGGUUUGUAAUGU-GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUUGUGGUUUACCCAGAGAGCGAAAACG ------.............-----...((((((--(.(((((.(..(((...-.)))..)))))).....(((..((((((........))))))....))).))))))).... ( -27.10, z-score = -2.57, R) >droSim1.chr2L 14159210 100 - 22036055 ------AUUUCCCUUUUUU-----CAGUUCGCU--UGUGUGGUUUGUAAUGU-GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUUGUGGUUUACCCGGAGAGCGAGAACG ------.............-----...((((((--(.(((((.(..(((...-.)))..)))))).....(((..((((((........))))))....))).))))))).... ( -26.10, z-score = -1.76, R) >droSec1.super_3 671046 100 - 7220098 ------AUUUCCCUUUUUU-----CAGUUCGCU--UGUGUGGUUUGUAAUGU-GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUUGUGGUUUACCCGGAGAGCGAGAACG ------.............-----...((((((--(.(((((.(..(((...-.)))..)))))).....(((..((((((........))))))....))).))))))).... ( -26.10, z-score = -1.76, R) >droYak2.chr2L 14516093 89 - 22324452 ------AUUUCCUUUUUU------CAGUCCGCU--UGUGGGUUUUGUAAUGU-GUUGUGGCCACAAAAU----------ACUUAAUUUUGCGGUUCACCCAGAGAGCGGGAGCG ------............------...((((((--(.(((((.((((((.((-(((.((....)).)))----------))......))))))...)))))..))))))).... ( -24.70, z-score = -1.31, R) >droEre2.scaffold_4929 6164923 105 - 26641161 ------AUUUCCUCUUUUUUUCAACAGUUCGCU--UGUGGGGUUUGUAAUGU-GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUUGUGGUUUGCCCAGAGAGCGAGAACG ------.((((((((((.....((((((....(--(((((...(..(((...-.)))..)))))))...))))))((((((........)))))).....)))))).))))... ( -28.00, z-score = -1.34, R) >droAna3.scaffold_12916 14773033 104 + 16180835 UUUGGUUUUUGUGUGUUU------UCGUUUGUU-UU-UGUGGUUUGUAAUGC-GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUGGUGGUUAACCCCGGAGCGAGAAGU- ...............(((------(((((((((-((-(((((.(..(((...-.)))..))))))))).)).((((((((((......))))))))))....)))))))))..- ( -32.90, z-score = -3.40, R) >dp4.chr4_group1 1778161 105 + 5278887 -UUAUUUUUUUUUUUUGC------CAGUCAGCU--UUUGUGGUUUGUAAUGCGGUUGUGGCCACAAAAUACUGUUGGCCACUUAAUUUUGUGGUUAACACGAGGAUAGAGACUG -.................------(((((...(--(((((((.(..((((...))))..)))))))))...((((((((((........))))))))))..........))))) ( -30.60, z-score = -2.57, R) >droPer1.super_5 1722124 99 - 6813705 -------UUUUUUUUUGC------CAGCCAGCU--UUUGUGGUUUGUAAUGCGGUUGUGGCCACAAAAUACUGUUGGCCACUUAAUUUUGUGGUUAACACGAGGAUAGAGACUG -------.......((((------.(((((.(.--...)))))).))))..((((((((((((((......)).)))))))....(((((((.....))))))).....))))) ( -28.70, z-score = -1.67, R) >droWil1.scaffold_180764 2669439 107 - 3949147 ------UUUCCCGUUUUUUCCUCCUACUUUUCUGGGUUGCUAUUUGUUUCUGCGGUUU-UUCGUAAUGUGUUUGUGGCCACUUAAUUCUGUGGUUAACACAAAAAUGUCCCACA ------..........................((((....(((((.....(((((...-.))))).(((((...(((((((........)))))))))))).))))).)))).. ( -21.60, z-score = -1.70, R) >droGri2.scaffold_15126 721406 89 - 8399593 ------GUUUGUGUGUCU------GAGUGUGCU-CUGUGUGUUUUGUG-------UGUGGCCGC-AAAUUCAGUUAUCCACUUAAUUUUGUGGUUAACAAACACACAAAU---- ------((((((((((.(------((((.(((.-(((..(.......)-------..)))..))-).)))))((((.((((........)))).))))..))))))))))---- ( -27.40, z-score = -2.82, R) >consensus ______AUUUCCUUUUUU______CAGUUCGCU__UGUGUGGUUUGUAAUGU_GUUGUGGCCACAAAAUACUGUUAGCCACUUAAUUUUGUGGUUAACCCAGAGAGCGAGAACG .......((((.((((.....................(((((.(..((((...))))..)))))).......((.((((((........)))))).))....)))).))))... (-10.46 = -11.36 + 0.90)
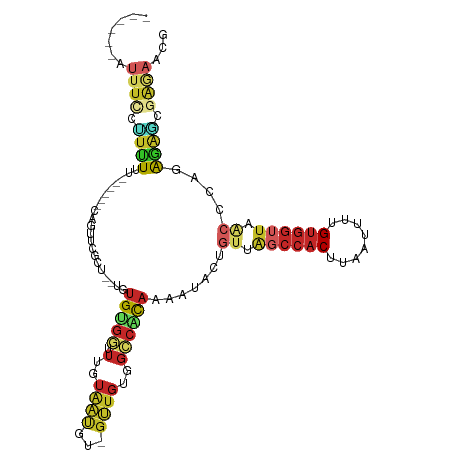
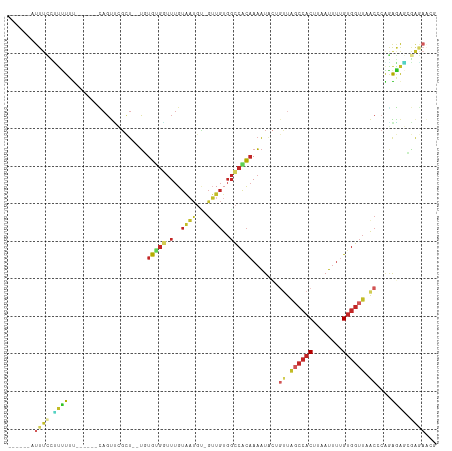
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:03 2011