
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,409,762 – 14,409,857 |
| Length | 95 |
| Max. P | 0.708459 |

| Location | 14,409,762 – 14,409,857 |
|---|---|
| Length | 95 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 70.50 |
| Shannon entropy | 0.61086 |
| G+C content | 0.54559 |
| Mean single sequence MFE | -25.25 |
| Consensus MFE | -13.34 |
| Energy contribution | -13.15 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.47 |
| Mean z-score | -0.82 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.708459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
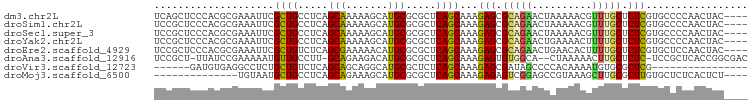
>dm3.chr2L 14409762 95 - 23011544 UCAGCUCCCACGCGAAAUUCGCUGCCUCAGCAAAAAGCAUGCGCGCUCAGCAAAGAGCGCAGAACUAAAAACGUUUGCUCUCGUGCCCCAACUAC---- ..........(((((.....((((...))))....(((....((((((......)))))).((((.......))))))).)))))..........---- ( -24.00, z-score = -0.85, R) >droSim1.chr2L 14157398 95 - 22036055 UCCGCUCCCACGCGAAAUUCGCUGCCUCAGCAAAAAGCAUGCGCGCUCAGCAAAGAGCGCAGAACUAAAAACGUUUGCUCUCGUGCCCCAACUAC---- ..(((......)))......((((...)))).....(((((.((((((......)))))).((((.......)))).....))))).........---- ( -24.50, z-score = -1.16, R) >droSec1.super_3 669230 95 - 7220098 UCCGCUCCCACGCGAAAUUCGCUGCCUCAGCAAAAAGCAUGCGCGCUCAGCAAAGAGCGCAGAACUAAAAACGUUUGCUCUCGUGCCCCAACUAC---- ..(((......)))......((((...)))).....(((((.((((((......)))))).((((.......)))).....))))).........---- ( -24.50, z-score = -1.16, R) >droYak2.chr2L 14514288 95 - 22324452 UCCGCUCCCACGCGAAAUUCGCUGCCUCAGCAAAAAGCAUGCGCGCUCAGCAAAGAGCGCAGAACUGAAAACUUUUGCUCUCGUGCCCCAACUAC---- ...(((.....(((.....)))(((....)))...)))..((((((((......))))((((((.(....).))))))....)))).........---- ( -24.80, z-score = -1.06, R) >droEre2.scaffold_4929 6163166 95 - 26641161 UCCGCUCCCACGCGAAAUUCGCUGUCUCAGCGAAAAACAUGCGCGCUCAGCAAAGAGCGCAGAACUGAACACUUUUGCUCUCGUGCUCCAACUAC---- ..(((......)))...(((((((...)))))))......((((((((......))))((((((........))))))....)))).........---- ( -27.70, z-score = -1.96, R) >droAna3.scaffold_12916 14771368 94 + 16180835 UCCGCU-UUAUCCGAAAAAUGUUGCCUU-GCAGAAGACAUGCGCGCUCAGCAAAGAGUGUGGCA--CUAAAAACUUGCUCUC-UCCGCUCACCGGCGAC ......-.............((((((..-((.(((((....(((((((......)))))))(((--.........)))))).-)).)).....)))))) ( -21.50, z-score = 0.53, R) >droVir3.scaffold_12723 3019327 77 + 5802038 ------GAUGUGAGGCCUCUGCUGUCUCAGCAGCAGGCAUGCGCUCUCAGCAAAGAGCGAUAGCCCCACAAAAUGUGCGCUCG---------------- ------..((((.(((.((((((((....))))))))....((((((......))))))...))).)))).............---------------- ( -28.80, z-score = -1.21, R) >droMoj3.scaffold_6500 16314420 81 + 32352404 --------------UGUAAUGCUGCCUCAGCAGAAAGCAUGCGCGCUCAGCAAAGAGAGUCGGAGCCGUAAAGCUUGCGCUUGUGCUCUCACUCU---- --------------.....(((((.....(((.......))).....)))))...(((((.(((((....((((....))))..))))).)))))---- ( -26.20, z-score = 0.31, R) >consensus UCCGCUCCCACGCGAAAUUCGCUGCCUCAGCAAAAAGCAUGCGCGCUCAGCAAAGAGCGCAGAACUAAAAACGUUUGCUCUCGUGCCCCAACUAC____ ....................((((.....(((.......))).....))))...(((.(((((..........))))).)))................. (-13.34 = -13.15 + -0.19)

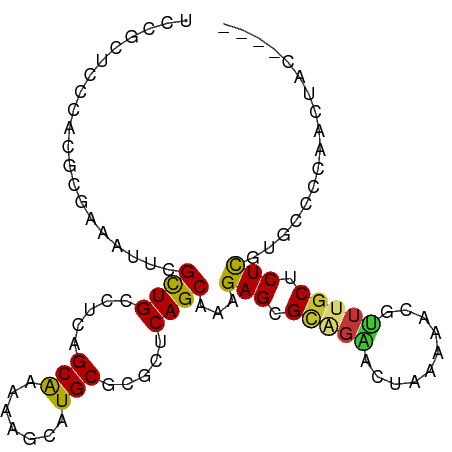

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:40:02 2011