
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 14,398,065 – 14,398,170 |
| Length | 105 |
| Max. P | 0.744398 |

| Location | 14,398,065 – 14,398,170 |
|---|---|
| Length | 105 |
| Sequences | 11 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 80.77 |
| Shannon entropy | 0.38247 |
| G+C content | 0.46083 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -15.51 |
| Energy contribution | -15.28 |
| Covariance contribution | -0.23 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.744398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14398065 105 + 23011544 ---GAGAGAAACACAAAACCCUGUCACAUUAACUGUCAACUGCCGCACA--ACUGACACUUGUCAUAUUGACAACCGACAGCGUCAUUUGACUGUCA-----AACGCGGGCCAAC ---...............(((((((........(((((..((.....))--..))))).(((((.....)))))..))))((((...(((.....))-----))))))))..... ( -22.00, z-score = -0.79, R) >droWil1.scaffold_180764 2647244 98 + 3949147 ----------ACACAAAACGCUGUCACAUUAACUGUCAACUGCUGCAAA--ACUGACACUUGUCAUAUUGACAAGUGACAGCGUCAUUUGACUGUCA-----AACUCAGACCAAA ----------...((((((((((((((......(((((...........--..))))).(((((.....)))))))))))))))..)))).(((...-----....)))...... ( -28.32, z-score = -3.30, R) >dp4.chr4_group1 1759914 115 - 5278887 AGAGAGAGAAACACAAAACGCUGUCACAUUAACUGUCAACUGCUGCAAAGCACUGACACUUGUCAUAUUGACAAGUGACAGCGUCAUUUGACUGUCAGAAAGAACAGAGACAGAC .............((((((((((((((......(((((..((((....)))).))))).(((((.....)))))))))))))))..)))).(((((............))))).. ( -38.10, z-score = -3.65, R) >droAna3.scaffold_12916 14761000 105 - 16180835 ---AAGAGAAACACAAAACGCUGUCACAUUAACUGUCAACUACUGCAGA--ACUGACACUUGUCAUAUUGACAAGUGACAGCGUCAUUUGACUGUCA-----AACCCGGGCCAAG ---....((....((((((((((((((.....((((........)))).--........(((((.....)))))))))))))))..))))....)).-----............. ( -27.80, z-score = -2.15, R) >droEre2.scaffold_4929 6151316 105 + 26641161 ---GGGAGAAACACAAAACCCUGUCACAUUAACUGUCAACUGCCGCACA--ACUGACACUUGUCAUAUUGACAACCGACAGCGUCAUUUGACUGUCA-----AACGCGGGCCAAC ---...............(((((((........(((((..((.....))--..))))).(((((.....)))))..))))((((...(((.....))-----))))))))..... ( -22.00, z-score = -0.26, R) >droYak2.chr2L 14502452 105 + 22324452 ---GGGAGAAACACAAAACCCUGUCACAUUAACUGUCAACUGCCGCACA--ACUGACACUUGUCAUAUUGACAACCGACAGCGUCAUUUGACUGUCA-----AACGCGGGCCAAC ---...............(((((((........(((((..((.....))--..))))).(((((.....)))))..))))((((...(((.....))-----))))))))..... ( -22.00, z-score = -0.26, R) >droSec1.super_3 657686 104 + 7220098 ---GAGAGAA-CACAAAACCCUGUCACAUUAACUGUCAACUGCCGCACA--ACUGACACUUGUCAUAUUGACAACCGACAGCGUCAUUUGACUGUCA-----AACGCGGGCCAAC ---.......-.......(((((((........(((((..((.....))--..))))).(((((.....)))))..))))((((...(((.....))-----))))))))..... ( -22.00, z-score = -0.74, R) >droSim1.chr2L 14145870 105 + 22036055 ---GAGAGAAACACAAAACCCUGUCACAUUAACUGUCAACUGCCGCACA--ACUGACACUUGUCAUAUUGACAACCGACAGCGUCAUUUGACUGUCA-----AACGCGGGCCAAC ---...............(((((((........(((((..((.....))--..))))).(((((.....)))))..))))((((...(((.....))-----))))))))..... ( -22.00, z-score = -0.79, R) >droGri2.scaffold_15126 703369 87 + 8399593 ------------------CGCUGUCACAUUAACUGUCAACUGCUGCAAA--ACUGACACUUGUCAUAUUGACAGCUGACAGCGUCAUUUGACUGUCAAACUUGGCCC-------- ------------------.(((((((((.....))).....(((((((.--..((((....))))..))).)))).))))))(((....))).((((....))))..-------- ( -24.70, z-score = -2.44, R) >droVir3.scaffold_12723 3007746 87 - 5802038 ------------------CGCUGUCACAUUAACUGUCAACUGCUGCAAA--AGUGACACUUGUCAUAUUGACAACUGACAGCGUCAUUUGACUGUCAAACUCGGCUC-------- ------------------.(((((((.......(((((.((........--))))))).(((((.....))))).)))))))(((....))).(((......)))..-------- ( -24.50, z-score = -2.18, R) >droMoj3.scaffold_6500 16298224 95 - 32352404 ------------------CGCUGUCACAUUAACUGUCAACUGCUGCAAA--AGUGACACUUGUCAUAUUGACAGCUGACAGCGUCAUUUGGCUGUCAAAGGCAGCUCGGCUCUAC ------------------.((((((................(((((((.--.(((((....))))).))).))))(((((((........)))))))..)))))).......... ( -32.90, z-score = -2.59, R) >consensus ___GAGAGAAACACAAAACGCUGUCACAUUAACUGUCAACUGCUGCAAA__ACUGACACUUGUCAUAUUGACAACUGACAGCGUCAUUUGACUGUCA_____AACGCGGGCCAAC ....................(((((........(((((...............))))).(((((.....)))))..))))).(((....)))....................... (-15.51 = -15.28 + -0.23)


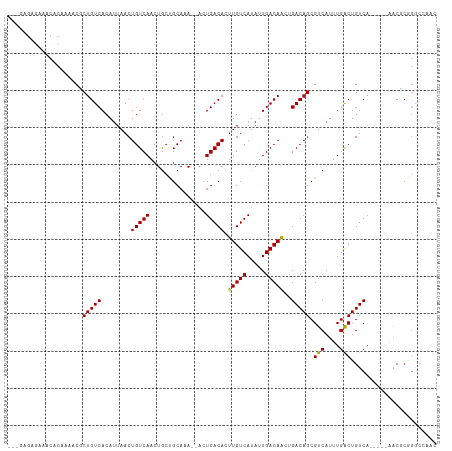
| Location | 14,398,065 – 14,398,170 |
|---|---|
| Length | 105 |
| Sequences | 11 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 80.77 |
| Shannon entropy | 0.38247 |
| G+C content | 0.46083 |
| Mean single sequence MFE | -31.25 |
| Consensus MFE | -19.71 |
| Energy contribution | -19.41 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.542586 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 14398065 105 - 23011544 GUUGGCCCGCGUU-----UGACAGUCAAAUGACGCUGUCGGUUGUCAAUAUGACAAGUGUCAGU--UGUGCGGCAGUUGACAGUUAAUGUGACAGGGUUUUGUGUUUCUCUC--- ...(((((..(((-----..((((((....)))((((((..(((((.(((((((....))))..--)))..)))))..))))))...)))))).))))).............--- ( -31.10, z-score = -0.82, R) >droWil1.scaffold_180764 2647244 98 - 3949147 UUUGGUCUGAGUU-----UGACAGUCAAAUGACGCUGUCACUUGUCAAUAUGACAAGUGUCAGU--UUUGCAGCAGUUGACAGUUAAUGUGACAGCGUUUUGUGU---------- ....(((......-----.)))...((...(((((((((((((((((((.((((....))))((--......)).)))))))).....)))))))))))...)).---------- ( -32.20, z-score = -2.02, R) >dp4.chr4_group1 1759914 115 + 5278887 GUCUGUCUCUGUUCUUUCUGACAGUCAAAUGACGCUGUCACUUGUCAAUAUGACAAGUGUCAGUGCUUUGCAGCAGUUGACAGUUAAUGUGACAGCGUUUUGUGUUUCUCUCUCU (.(((((............))))).)....((((((((((((((((.....))))).((((((((((....)))).))))))......)))))))))))................ ( -39.10, z-score = -3.25, R) >droAna3.scaffold_12916 14761000 105 + 16180835 CUUGGCCCGGGUU-----UGACAGUCAAAUGACGCUGUCACUUGUCAAUAUGACAAGUGUCAGU--UCUGCAGUAGUUGACAGUUAAUGUGACAGCGUUUUGUGUUUCUCUU--- ........(((..-----.((((..(((..(((((((((((((((((((.((((....))))(.--....)....)))))))).....)))))))))))))))))).)))..--- ( -32.10, z-score = -1.70, R) >droEre2.scaffold_4929 6151316 105 - 26641161 GUUGGCCCGCGUU-----UGACAGUCAAAUGACGCUGUCGGUUGUCAAUAUGACAAGUGUCAGU--UGUGCGGCAGUUGACAGUUAAUGUGACAGGGUUUUGUGUUUCUCCC--- ...(((((..(((-----..((((((....)))((((((..(((((.(((((((....))))..--)))..)))))..))))))...)))))).))))).............--- ( -31.10, z-score = -0.79, R) >droYak2.chr2L 14502452 105 - 22324452 GUUGGCCCGCGUU-----UGACAGUCAAAUGACGCUGUCGGUUGUCAAUAUGACAAGUGUCAGU--UGUGCGGCAGUUGACAGUUAAUGUGACAGGGUUUUGUGUUUCUCCC--- ...(((((..(((-----..((((((....)))((((((..(((((.(((((((....))))..--)))..)))))..))))))...)))))).))))).............--- ( -31.10, z-score = -0.79, R) >droSec1.super_3 657686 104 - 7220098 GUUGGCCCGCGUU-----UGACAGUCAAAUGACGCUGUCGGUUGUCAAUAUGACAAGUGUCAGU--UGUGCGGCAGUUGACAGUUAAUGUGACAGGGUUUUGUG-UUCUCUC--- ...(((((..(((-----..((((((....)))((((((..(((((.(((((((....))))..--)))..)))))..))))))...)))))).))))).....-.......--- ( -31.10, z-score = -0.79, R) >droSim1.chr2L 14145870 105 - 22036055 GUUGGCCCGCGUU-----UGACAGUCAAAUGACGCUGUCGGUUGUCAAUAUGACAAGUGUCAGU--UGUGCGGCAGUUGACAGUUAAUGUGACAGGGUUUUGUGUUUCUCUC--- ...(((((..(((-----..((((((....)))((((((..(((((.(((((((....))))..--)))..)))))..))))))...)))))).))))).............--- ( -31.10, z-score = -0.82, R) >droGri2.scaffold_15126 703369 87 - 8399593 --------GGGCCAAGUUUGACAGUCAAAUGACGCUGUCAGCUGUCAAUAUGACAAGUGUCAGU--UUUGCAGCAGUUGACAGUUAAUGUGACAGCG------------------ --------...............((((.((...(((((((((((((((..((((....))))..--.)))..))))))))))))..)).))))....------------------ ( -29.30, z-score = -2.13, R) >droVir3.scaffold_12723 3007746 87 + 5802038 --------GAGCCGAGUUUGACAGUCAAAUGACGCUGUCAGUUGUCAAUAUGACAAGUGUCACU--UUUGCAGCAGUUGACAGUUAAUGUGACAGCG------------------ --------.......(((((.....)))))..(((((((.(((((.((..((((....))))..--)).))))).....(((.....))))))))))------------------ ( -25.20, z-score = -0.91, R) >droMoj3.scaffold_6500 16298224 95 + 32352404 GUAGAGCCGAGCUGCCUUUGACAGCCAAAUGACGCUGUCAGCUGUCAAUAUGACAAGUGUCACU--UUUGCAGCAGUUGACAGUUAAUGUGACAGCG------------------ ((((.......))))...(((((((........)))))))(((((((...((((...(((((((--........)).)))))))))...))))))).------------------ ( -30.40, z-score = -1.38, R) >consensus GUUGGCCCGCGUU_____UGACAGUCAAAUGACGCUGUCAGUUGUCAAUAUGACAAGUGUCAGU__UGUGCAGCAGUUGACAGUUAAUGUGACAGCGUUUUGUGUUUCUCUC___ ..................(((((((........))))))).((((((...((((...((((((.............))))))))))...)))))).................... (-19.71 = -19.41 + -0.30)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:39:58 2011