
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,275,066 – 1,275,167 |
| Length | 101 |
| Max. P | 0.593697 |

| Location | 1,275,066 – 1,275,167 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 69.73 |
| Shannon entropy | 0.58900 |
| G+C content | 0.56958 |
| Mean single sequence MFE | -20.29 |
| Consensus MFE | -9.89 |
| Energy contribution | -10.02 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.97 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.593697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 1275066 101 - 23011544 AAUUCUCUAUCCCGA-AUCCCAGCUCCAUAUAGCCCAAAUCCCCUUCCCACGCAGUGGAGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC ...........((((-(((.(((((((((...((.................)).))))).)))).)))))....(((.......))).......--...))... ( -19.03, z-score = -0.69, R) >droSim1.chr2L 1247739 101 - 22036055 AAUCCUCUAUCCCGA-AUCCCCGCUCCAUCCAGCCCAAAUCCCCAUCCCACGCAGCGGAGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC ...............-....((((..........................((((((....))))))........(((.......))).......--.))))... ( -19.00, z-score = -0.57, R) >droSec1.super_14 1225586 101 - 2068291 AAUCCUCUAUCCCGA-GUCCCCGCUCCAUCCAGGCCAAGUCCCCAUCCCACGCAGCGGAGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC .............((-((....))))......((((..............((((((....))))))........(((.......))).......--...)))). ( -21.00, z-score = 0.02, R) >droYak2.chr2L 1250848 102 - 22324452 AAUCCUCUAUCCUGGUGUCCCCGCACCAUUGAACCCGAAUGCCCAUCCCACGCAGUCGAGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC ....(((.....((((((....))))))........((.(((.........))).)))))((((((........(((.......))).......--)))))).. ( -23.46, z-score = -0.77, R) >droEre2.scaffold_4929 1315793 97 - 26641161 AAUCCUCCUUCCCGA-----GUGCCGCACUCAACCCGAAUCCCCAUCCCACGCAGUCGUGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC ....(((......))-----).(((((.......................((((((....))))))........(((.......))).......--.))))).. ( -23.50, z-score = -2.20, R) >droAna3.scaffold_12916 4737577 79 - 16180835 -------------------GGUCCUU---CGCAGUUGCCACCAGCUACC-CACAAUUGAGACUGCGAUUUCAACUGCAUCAUCAGCAUAAUCAA--CGCGGCUC -------------------((((...---.(((((((...((((((...-........)).))).)....))))))).......((........--.)))))). ( -14.10, z-score = 0.36, R) >dp4.chr4_group3 9748320 93 - 11692001 ---------UCCCC--UCCCGCUUUCACCCCCGCCCACCAGCAGCAGUCGCCCAAUUGAGACUGCGAUUCCGAGUGCAUCAUCAGCAUAAUCAACGCGCGGCUC ---------.....--..((((..........((......)).((((((..........))))))((((....((((.......)))))))).....))))... ( -21.40, z-score = -1.44, R) >droPer1.super_8 947471 89 - 3966273 ---------UCCCC--ACCCGCUUU----CCCACCCACCACCAGCAGUCGCCCAAUUGAGACUGCGAUUCCGAGUGCAUCAUCAGCAUAAUCAACGCGCGGCUC ---------.....--..((((...----..............((((((..........))))))((((....((((.......)))))))).....))))... ( -20.80, z-score = -2.46, R) >consensus AAUCCUCUAUCCCGA_AUCCCCGCUCCAUCCAACCCAAAUCCCCAUCCCACGCAGUGGAGGCUGCGAUUCCAACUGCAUCAUCAGCAUAAUCAA__CGCGGCUC .........................................................(((.(((((........(((.......))).........)))))))) ( -9.89 = -10.02 + 0.13)

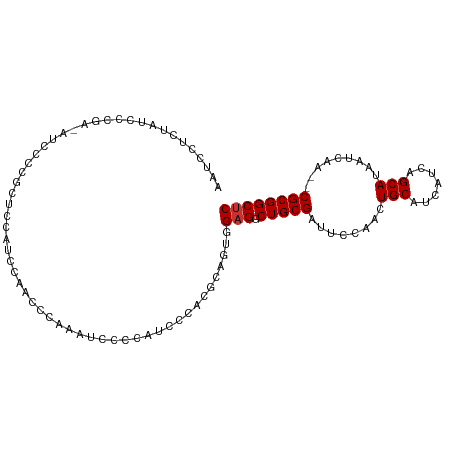

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:09 2011