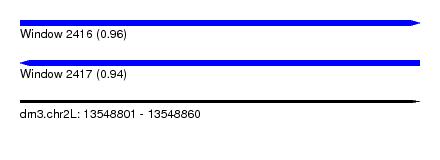

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 13,548,801 – 13,548,860 |
| Length | 59 |
| Max. P | 0.960146 |

| Location | 13,548,801 – 13,548,860 |
|---|---|
| Length | 59 |
| Sequences | 6 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 86.40 |
| Shannon entropy | 0.26050 |
| G+C content | 0.52814 |
| Mean single sequence MFE | -22.83 |
| Consensus MFE | -18.76 |
| Energy contribution | -18.96 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.960146 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
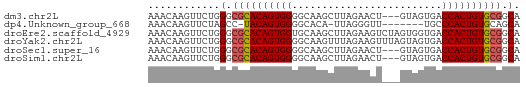
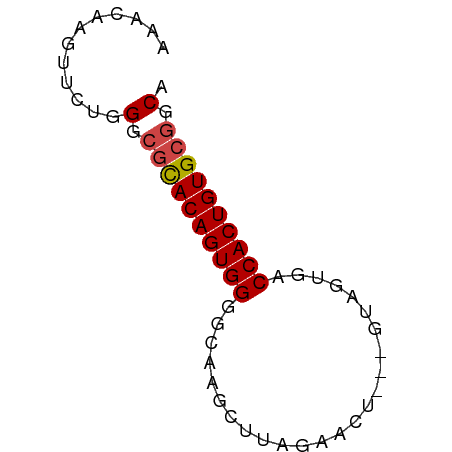
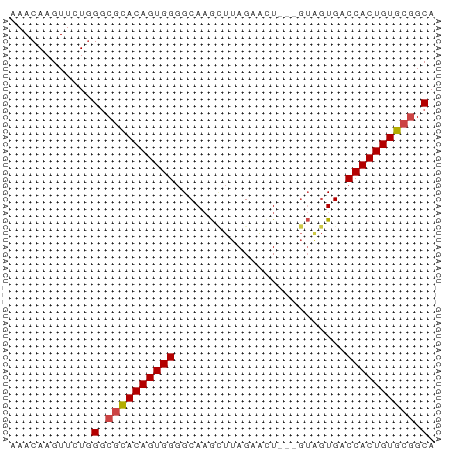
>dm3.chr2L 13548801 59 + 23011544 AAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCA ............(.((((((((((..((.((........---))..)).)))))))))).). ( -23.20, z-score = -2.07, R) >dp4.Unknown_group_668 8459 53 - 12025 AAACAAGUUCUAGCC-UACAGUGGGGCACA-UUAGGGUU-------UGCCCACUGUGCAGCA ............((.-(((((((((.((.(-(....)).-------)))))))))))..)). ( -17.00, z-score = -0.73, R) >droEre2.scaffold_4929 14738399 62 - 26641161 AAACAAGUUCUGGGCGCACAGUGGUGCAAGCUUAGAAGUCUAGUGGUGACCACUGUGCGGCA ............(.(((((((((((.((.(((.((....)))))..))))))))))))).). ( -27.40, z-score = -3.13, R) >droYak2.chr2L 9973012 62 + 22324452 AAACAAGUUCUGGGCGCACAGUGGGGCAAGUUUAGAAGUUUAGUAGUGACCACUGUGCGGCA ............(.((((((((((..((...((((....))))...)).)))))))))).). ( -23.00, z-score = -2.84, R) >droSec1.super_16 1688671 59 + 1878335 AAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCA ............(.((((((((((..((.((........---))..)).)))))))))).). ( -23.20, z-score = -2.07, R) >droSim1.chr2L 13321949 59 + 22036055 AAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCA ............(.((((((((((..((.((........---))..)).)))))))))).). ( -23.20, z-score = -2.07, R) >consensus AAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU___GUAGUGACCACUGUGCGGCA ............(.((((((((((.........................)))))))))).). (-18.76 = -18.96 + 0.20)
| Location | 13,548,801 – 13,548,860 |
|---|---|
| Length | 59 |
| Sequences | 6 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 86.40 |
| Shannon entropy | 0.26050 |
| G+C content | 0.52814 |
| Mean single sequence MFE | -17.98 |
| Consensus MFE | -15.02 |
| Energy contribution | -15.21 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.936641 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
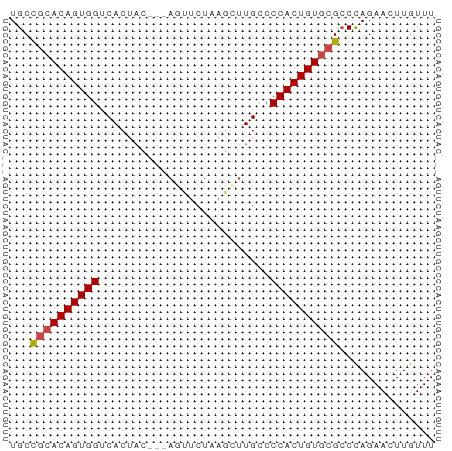
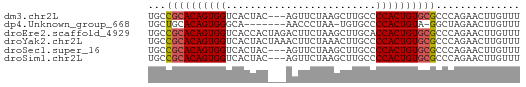
>dm3.chr2L 13548801 59 - 23011544 UGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUU ...((((((((((.((....---(((.....)))))..)))))))))).............. ( -17.50, z-score = -1.61, R) >dp4.Unknown_group_668 8459 53 + 12025 UGCUGCACAGUGGGCA-------AACCCUAA-UGUGCCCCACUGUA-GGCUAGAACUUGUUU .(((..((((((((..-------.((.....-.))..)))))))).-)))............ ( -17.60, z-score = -1.45, R) >droEre2.scaffold_4929 14738399 62 + 26641161 UGCCGCACAGUGGUCACCACUAGACUUCUAAGCUUGCACCACUGUGCGCCCAGAACUUGUUU ...(((((((((((((....(((....)))....)).))))))))))).............. ( -21.70, z-score = -2.99, R) >droYak2.chr2L 9973012 62 - 22324452 UGCCGCACAGUGGUCACUACUAAACUUCUAAACUUGCCCCACUGUGCGCCCAGAACUUGUUU ...((((((((((.........................)))))))))).............. ( -16.11, z-score = -2.38, R) >droSec1.super_16 1688671 59 - 1878335 UGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUU ...((((((((((.((....---(((.....)))))..)))))))))).............. ( -17.50, z-score = -1.61, R) >droSim1.chr2L 13321949 59 - 22036055 UGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUU ...((((((((((.((....---(((.....)))))..)))))))))).............. ( -17.50, z-score = -1.61, R) >consensus UGCCGCACAGUGGUCACUAC___AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUU ...((((((((((.........................)))))))))).............. (-15.02 = -15.21 + 0.19)
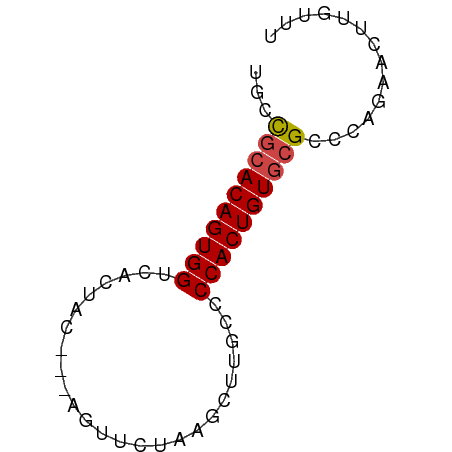

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:38:15 2011