
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 13,498,691 – 13,498,790 |
| Length | 99 |
| Max. P | 0.839865 |

| Location | 13,498,691 – 13,498,790 |
|---|---|
| Length | 99 |
| Sequences | 12 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 65.35 |
| Shannon entropy | 0.72078 |
| G+C content | 0.64019 |
| Mean single sequence MFE | -40.93 |
| Consensus MFE | -14.95 |
| Energy contribution | -15.00 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.78 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.839865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
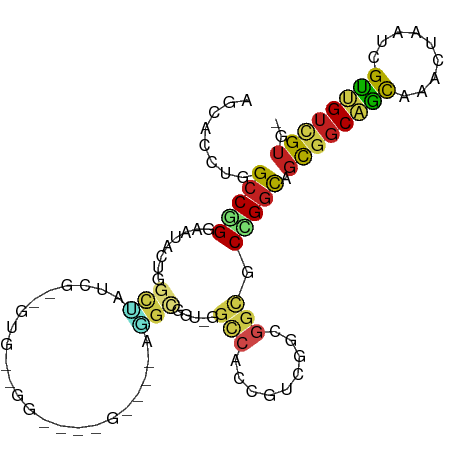
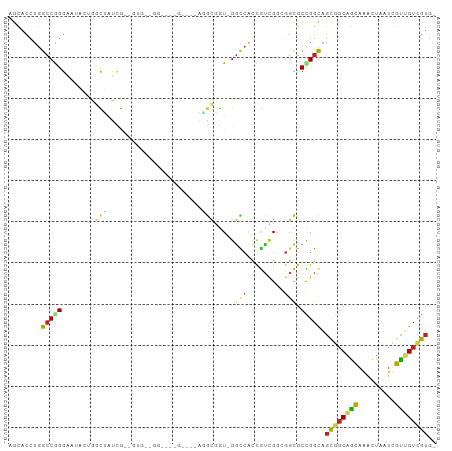
>dm3.chr2L 13498691 99 - 23011544 AGCACCUGGCCGGGAAUACUGGCUAUCGUUGUGCUGG---UGGAUGAGGCGGU-GGCCACCGUCGAAGGCGCCGGCAGCGGCAGCAGACUAAUCGCUGUCGUG- (((...(((((((.....)))))))..))).((((((---((..(..((((((-....))))))..)..))))))))((((((((.((....)))))))))).- ( -46.60, z-score = -1.79, R) >droSim1.chr2L 13280592 99 - 22036055 AGCACCUGGCCGGGAAUACUGGCUAUCGUUGUGCUGG---UGGAUGAGGCGGU-GGCCACCGUCGAAGGCGCCGGCAGCGGCAGCAAACUAAUCGCUGUCGUG- (((...(((((((.....)))))))..))).((((((---((..(..((((((-....))))))..)..))))))))((((((((.........)))))))).- ( -46.20, z-score = -1.93, R) >droSec1.super_16 1644013 99 - 1878335 AGCACCUGGCCGGGAAUACAGGCUAUCGUUGUGCUGG---UGGAUGAGGCGGU-GGCCACCGUCGAAGGCGCCGGCAGCGGCAGCAAACUAAUCGUUGUCGUG- (((...(((((.(.....).)))))..))).((((((---((..(..((((((-....))))))..)..))))))))((((((((.........)))))))).- ( -39.90, z-score = -0.48, R) >droYak2.chr2L 9927040 99 - 22324452 AGCACCUGGCCGGGAAUACUGGCUAUCGUUGUGCUGG---UGGAUGAGGCGGU-GGCCACCGUCGGAGGCGCCGGCAGCGGCAGCAAACUAAUUGCUGUCGUG- (((...(((((((.....)))))))..))).((((((---((..(..((((((-....))))))..)..))))))))((((((((((.....)))))))))).- ( -48.40, z-score = -2.48, R) >droEre2.scaffold_4929 14690359 99 + 26641161 AGCACCUGGCCGGGAAUACUGGCUAUCGUUGUGCUGG---UGGAUGAGGCGGU-GGCCACCGUCGAAGGCGCCGGCAGCGGCAGCAAGCUAAUUGCUGUCGUG- (((...(((((((.....)))))))..))).((((((---((..(..((((((-....))))))..)..))))))))((((((((((.....)))))))))).- ( -49.60, z-score = -2.69, R) >droAna3.scaffold_12943 2203298 84 - 5039921 AGGACCUGGCCGGGAAUGCUGGCCACAG------------------GGGCGGU-GGCCAUUGUCGGCGGCGCCGGCAGCGGCAGCAAACUGAUCGUUGUGGUG- ...(((.((((((.....))))))((((------------------.((((((-.(((.((((((((...)))))))).))).....)))).)).))))))).- ( -39.20, z-score = -0.83, R) >dp4.chr4_group1 3224909 81 + 5278887 GGCACCUGGCCAGGAA---UACUGGUCA------------------UCGCCGU-GGCCAUUGUCGGCGGCGCCGGUAGCGGCAGCAGGCUGAUCGUUGUCGUG- (((...(((((((...---..)))))))------------------..(((((-(((....))).))))))))(..((((((((....))).)))))..)...- ( -37.70, z-score = -1.41, R) >droPer1.super_5 3174494 81 - 6813705 GGCACCUGGCCAGGAA---UACUGGUCA------------------UCGCCGU-GGCCAUUGUCGGCGGCGCCGGCAGCGGCAGCAGGCUGAUCGUUGUCGUG- (((...(((((((...---..)))))))------------------..(((((-(((....))).))))))))(((((((((((....))).))))))))...- ( -41.60, z-score = -2.30, R) >droWil1.scaffold_180708 7335417 81 + 12563649 GGUAUUUGGCCAGGAA---UGGCCAUUG------------------ACAUUGU-GGCCAUUGUGGGCGGCGCUGGCAGUGGCAACAGACUAAUCGUUGUGGUG- .((...((((((....---))))))...------------------)).((((-.((((((((.(((...))).)))))))).))))((((.......)))).- ( -32.30, z-score = -1.98, R) >droVir3.scaffold_12723 5332365 90 + 5802038 AGCACCUGGCCAGGUAUGCUCGCUCUUGCAGUCGACG---------UUGUAGU-G---GCCGUUGGCGGCGCCGGCAGCGGCAGUAAACUGAUCGUCGUUGUG- (((((((....))))..))).......((((.(((((---------...((((-.---(((((((.((....)).))))))).....))))..))))))))).- ( -36.40, z-score = -0.44, R) >droGri2.scaffold_15252 13131440 102 + 17193109 GGCACCUGGCCGGGUAUUAAGCUGCCUGCAGCGGAGGAUCCACUGUUUGCAAU-GCUCGCCAGUGGCGGCGCUGGCAGCGGCAGCAGACUAAUCGUCGUUGUG- (((((((....)))).....((((((((((((((.(....).))).)))))..-(((.(((((((....)))))))))))))))).........)))......- ( -43.70, z-score = -0.57, R) >anoGam1.chr3R 32112408 88 + 53272125 GGCAUGAAGCCGGGCA--GCGGCGAGGA------------AGUGGAUGACGAUCGGUAACCGCCGGC--UACCGACGACGACGGCAGGUACGACGUCGAUGGCA (((.....((((....--.))))..((.------------..(.(((....))).)...))))).((--((.(((((.((((.....)).)).))))).)))). ( -29.60, z-score = 0.38, R) >consensus AGCACCUGGCCGGGAAUACUGGCUAUCG__GUG__GG____G____AGGCGGU_GGCCACCGUCGGCGGCGCCGGCAGCGGCAGCAAACUAAUCGUUGUCGUG_ ........(((((...................................((.....))...(((.....)))))))).((((((((.........)))))))).. (-14.95 = -15.00 + 0.05)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:38:06 2011