| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,242,856 – 1,242,946 |
| Length | 90 |
| Max. P | 0.669212 |

| Location | 1,242,856 – 1,242,946 |
|---|---|
| Length | 90 |
| Sequences | 7 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 69.13 |
| Shannon entropy | 0.53157 |
| G+C content | 0.59091 |
| Mean single sequence MFE | -22.49 |
| Consensus MFE | -12.89 |
| Energy contribution | -12.64 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.669212 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
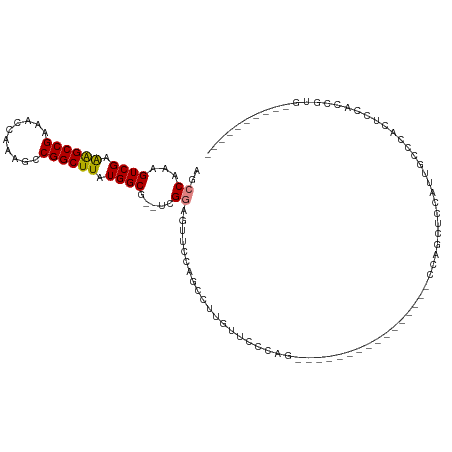
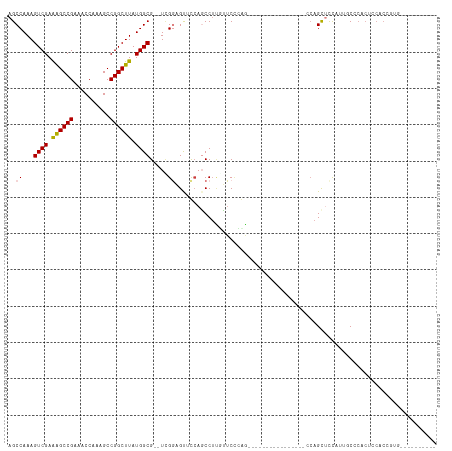
>dm3.chr2L 1242856 90 + 23011544 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UCGGAGUUCCAGCCUUCUUCUCGA----------------CCAGCUCCAUUGCCCACUCCACAGUG---------- (((....(((((((((((..........))))))..(((.--..((....)).)))......))))----------------)..)))........((((....))))---------- ( -22.20, z-score = -0.75, R) >droSim1.chr2L 1215068 90 + 22036055 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UUGGAGUUCCAGCCUUGUUCCCGG----------------CCAGCUCCAUUGCCCACUCCACGGUG---------- .(((........((((((..........))))))..((((--.(((((((...(((........))----------------).))))))).))))........))).---------- ( -30.60, z-score = -2.03, R) >droSec1.super_14 1193801 90 + 2068291 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UCGGAGCUCCAGCCUUGUUCCCGG----------------CCAGCUCCAUUGCCCACUCCACCGUG---------- ...((..((.((((((((..........))))))..((((--..((((((...(((........))----------------).))))))..))))...)).))..))---------- ( -28.60, z-score = -1.83, R) >droYak2.chr2L 1217844 106 + 22324452 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UCGGAGUUCCAGCCUUGUUCUCCAGUUCUCCACUGCUCAGCCAGCUCCAUGGCCCACUCCACAGUU---------- .((((..((((.((((((..........)))))).)))).--..((((((...((..((.....((((.....))))..)))).)))))).)))).............---------- ( -28.10, z-score = -0.89, R) >droAna3.scaffold_12916 4704821 77 + 16180835 AGCCAAAGUCGAGGGCCGAAACCAAAGCCGGCUUAUGGCGAGUGGGAGUCUAAGCCAAUCCCGGAA------------------AGUAUUCCGCC----------------------- .((((((((((..((......)).....)))))).))))..((((((..((...((......))..------------------))..)))))).----------------------- ( -22.50, z-score = -0.19, R) >dp4.chr4_group3 9701761 96 + 11692001 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UCGUCGUCUCAUCCUCUGUCCAAGU--------------ACCAGUUCCCCCCCCCCCCCCCCCCACCCUU------ .......((((.((((((..........)))))).)))).--.........................--------------...............................------ ( -12.70, z-score = -0.86, R) >droPer1.super_8 900234 102 + 3966273 AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG--UCGUCGUCUCAUCCUCUGUCCAAGU--------------ACCAGUCCCCCCCCCCUCCUCCCCCCUCCCCACCGUU .......((((.((((((..........)))))).)))).--.........................--------------..................................... ( -12.70, z-score = -0.44, R) >consensus AGCCAAAGUCGAAAGCCGAAACCAAAGCCGGCUUAUGGCG__UCGGAGUUCCAGCCUUGUUCCCAG________________CCAGCUCCAUUGCCCACUCCACCGUG__________ .......((((.((((((..........)))))).))))............................................................................... (-12.89 = -12.64 + -0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:01 2011