| Sequence ID | dm3.chr2L |
|---|---|
| Location | 13,164,406 – 13,164,472 |
| Length | 66 |
| Max. P | 0.539771 |

| Location | 13,164,406 – 13,164,472 |
|---|---|
| Length | 66 |
| Sequences | 13 |
| Columns | 68 |
| Reading direction | reverse |
| Mean pairwise identity | 72.35 |
| Shannon entropy | 0.62466 |
| G+C content | 0.53091 |
| Mean single sequence MFE | -16.89 |
| Consensus MFE | -9.50 |
| Energy contribution | -8.83 |
| Covariance contribution | -0.67 |
| Combinations/Pair | 1.73 |
| Mean z-score | -0.45 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.539771 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
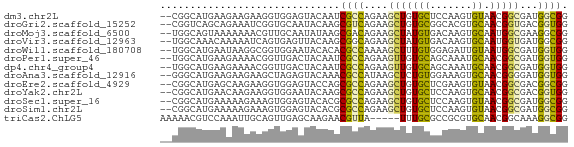
>dm3.chr2L 13164406 66 - 23011544 --CGGCAUGAAGAAGAAGGUGGAGUACAAUCGCCAGAAGCUGUGCUCCAAGUGUAACGGCGAUGGCGG --((.((((..........(((((((((...((.....)))))))))))..........).))).)). ( -18.35, z-score = -0.83, R) >droGri2.scaffold_15252 10193449 66 - 17193109 --CGGUCAGCAGAAAUCGGUGCAAUACAAGCGUCAGAAGCUGUGCGGCACGUGCAACGGUGACGGUGG --((((........))))((((..((((.((.......))))))..))))...((.((....)).)). ( -17.80, z-score = 0.60, R) >droMoj3.scaffold_6500 26230892 66 - 32352404 --UGGCAGUAAAAAAACGUUGCAAUAUAAGCGACAGAAGCUAUGUGACAAGUGCAAUGGCGAAGGCGG --..............(((((((.(((((((.......))).)))).....)))))))((....)).. ( -14.70, z-score = -1.03, R) >droVir3.scaffold_12963 7039782 66 - 20206255 --UGGCAAACAAAAAUCAGUGAGUUACAAGCGGCAGAAGCUAUGUGACAAGUGCAAUGGUGAUGGCGG --..((..((......((.(..(((((((((.......))).)))))).).)).....))....)).. ( -12.20, z-score = 0.38, R) >droWil1.scaffold_180708 6265088 66 - 12563649 --UGGCAUGAAUAAGGCGGUGGAAUACACACGCCAAAAGCUUUGUGGAGAUUGUAAUGGCGAUGGUGG --...(((.(....(((((((.....))).))))....((((((..(...)..))).)))..).))). ( -15.40, z-score = -1.02, R) >droPer1.super_46 411634 66 + 590583 --UGGCAUGAAGAAAACGGUUGACUACAAUCGCCAGAAGUUGUGCAGCAAAUGCAACGGCGAUGGUGG --...............(((((....)))))((((...((((((((.....))).)))))..)))).. ( -16.90, z-score = -0.78, R) >dp4.chr4_group4 882517 66 - 6586962 --UGGCAUGAAGAAAACGGUUGACUACAAUCGCCAGAAGUUGUGCAGCAAAUGCAACGGCGAUGGUGG --...............(((((....)))))((((...((((((((.....))).)))))..)))).. ( -16.90, z-score = -0.78, R) >droAna3.scaffold_12916 3899900 66 + 16180835 --GGGCAUGAAGAAGAAGCUAGAGUACAAACGCCAUAAGCUCUGUGGAAAGUGCAACGGGGAUGGUGG --.(((...........)))..........((((((...((((((..........)))))))))))). ( -13.70, z-score = -0.42, R) >droEre2.scaffold_4929 14365093 66 + 26641161 --CGGCAUGAGCAAGAAGGUGGAGUACCAGCGCCAGAAGCUGUGCUCGAAGUGUAACGGCGACGGCGG --..((....(((......(.((((((.(((.......))))))))).)..)))..((....)))).. ( -17.60, z-score = 0.80, R) >droYak2.chr2L 9587909 66 - 22324452 --CGGCAUGAACAAGAAGGUGGAAUACAAGCGCCAGAAGCUGUGCUCCAAGUGCAACGGCGACGGUGG --..((((...........((((.((((.((.......)))))).)))).))))..((....)).... ( -15.60, z-score = 0.71, R) >droSec1.super_16 1319226 66 - 1878335 --CGGCAUGAAAAAGAAAGUGGAGUACACGCGCCAGAAGCUGUGCUCCAAGUGUAACGGCGAUGGCGG --((.((((.......(..(((((((((.((.......)))))))))))..).......).))).)). ( -21.34, z-score = -1.62, R) >droSim1.chr2L 12959102 66 - 22036055 --CGGCAUGAAAAAGAAAGUGGAGUACACGCGCCAGAAGCUGUGCUCCAAGUGUAACGGCGAUGGCGG --((.((((.......(..(((((((((.((.......)))))))))))..).......).))).)). ( -21.34, z-score = -1.62, R) >triCas2.ChLG5 18200319 63 + 18847211 AAAAACGUCCAAAUUGCAGUUGAGCAAGAACGUUA-----UUUGCGCCGCGUGCAACGGCAAAGGCGG ...(((((.....((((......))))..))))).-----....((((...(((....)))..)))). ( -17.80, z-score = -0.25, R) >consensus __CGGCAUGAAAAAGACGGUGGAGUACAAGCGCCAGAAGCUGUGCGCCAAGUGCAACGGCGAUGGCGG ..............................((((....(((((((.......)).)))))...)))). ( -9.50 = -8.83 + -0.67)

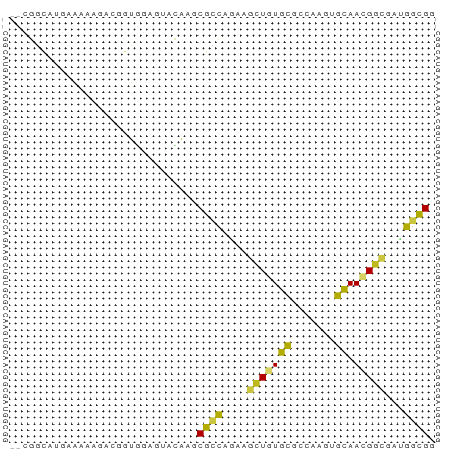
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:37:07 2011