
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,977,989 – 12,978,089 |
| Length | 100 |
| Max. P | 0.560425 |

| Location | 12,977,989 – 12,978,089 |
|---|---|
| Length | 100 |
| Sequences | 15 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 79.22 |
| Shannon entropy | 0.43197 |
| G+C content | 0.51984 |
| Mean single sequence MFE | -34.56 |
| Consensus MFE | -19.98 |
| Energy contribution | -19.40 |
| Covariance contribution | -0.58 |
| Combinations/Pair | 1.62 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.560425 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 12977989 100 + 23011544 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGAAAUCUGGAGGACUAGAGA-----CCGCGCUGGCCCUACUU------------- ..((((.(((((...(((((.....))).))((.((((((........)))))).(((((((((....))))))).)).....-----..))..))))).)))).------------- ( -36.40, z-score = -0.78, R) >droSim1.chr2L 12769753 100 + 22036055 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAGA-----CCGCGCUGGCCCUACUU------------- ..((((.(((((.......(((((........)))))((..((((.((((...)))).)))).((..(((((....)))))..-----))))..))))).)))).------------- ( -36.20, z-score = -0.39, R) >droSec1.super_16 1132507 100 + 1878335 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAGA-----CCGCGCUGGCCCUACUU------------- ..((((.(((((.......(((((........)))))((..((((.((((...)))).)))).((..(((((....)))))..-----))))..))))).)))).------------- ( -36.20, z-score = -0.39, R) >droYak2.chr2L 9396347 100 + 22324452 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAGA-----CCGCGUUGGCCCUACUU------------- ..((((.((((((......(((((........)))))((..((((.((((...)))).)))).((..(((((....)))))..-----)))).)))))).)))).------------- ( -37.20, z-score = -0.94, R) >droEre2.scaffold_4929 14172482 100 - 26641161 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAUA-----CCGCGUUGGCCCUACUU------------- ..((((.((((((......(((((........)))))((..((((.((((...)))).)))).((.((((((....)))))).-----)))).)))))).)))).------------- ( -38.10, z-score = -1.48, R) >droAna3.scaffold_12916 3717437 100 - 16180835 AUGGUGAGGCCAAGAUCAAGGCCGAUUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAGA-----CCGCAGUGGCCCUACUU------------- ..((((.((((.........((((....)).)).(((((..((((.((((...)))).)))).((..(((((....)))))..-----))))))))))).)))).------------- ( -38.50, z-score = -1.54, R) >droWil1.scaffold_180708 6008416 99 + 12563649 AUGGUGAGGCCAAGAUCAAGGCUGACUUUGAGCAGUUGCACGAAGAUUUGCAGCAAGCCUUCAGAAAUCUGGAGGACUAGAUC-----GCAU-UUGGUCUUACGU------------- ...(((((((((((.((((((....))))))((.((((((........)))))).(((((((((....))))))).)).....-----)).)-))))))))))..------------- ( -40.40, z-score = -4.18, R) >dp4.chr4_group4 645086 98 + 6586962 AUGGUGAGGCCAAGAUCAAGGCUGACUUUGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGAAAUCUGGAGGACUAGAG-------CGUGUUGGCCUUACUU------------- ..(((((((((((.(((((((....))))))((.((((((........)))))).(((((((((....))))))).))...)-------).).))))))))))).------------- ( -45.10, z-score = -3.93, R) >droPer1.super_46 178555 98 - 590583 AUGGUGAGGCCAAGAUCAAGGCUGACUUUGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGAAAUCUGGAGGACUAGAG-------CGUGUUGGCCUUACUU------------- ..(((((((((((.(((((((....))))))((.((((((........)))))).(((((((((....))))))).))...)-------).).))))))))))).------------- ( -45.10, z-score = -3.93, R) >droVir3.scaffold_12963 6826032 98 + 20206255 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCAUGAGGAUUUGCAGCAGGCCUUCAGGAAUCUGGAGGAUUAGAG-------CGUGUUGUCCGUACUU------------- ..((((.((((........))))(((..((.((.((((((........))))))...(((((((....)))))))......)-------).))..)))..)))).------------- ( -31.80, z-score = -0.40, R) >droMoj3.scaffold_6500 26007424 98 + 32352404 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAUCAGCUGCAUGAGGACUUGCAGCAGGCCUUCAGGAAUCUGGAGGAUUAAAG-------CGCGUUGUCCGUACUU------------- (((((((.(((..(((((((.....))).)))).((((((........))))))...(((((((....)))))))......)-------.)).))).))))....------------- ( -32.50, z-score = -0.71, R) >droGri2.scaffold_15252 10003442 98 + 17193109 AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCAUGAGGAUUUGCAGCAGGCCUUCAGAAAUCUGGAGGAUUAGAG-------CACGUUGUCCGUACUU------------- ..((((.((((........))))(((..((.((.((((((........))))))...(((((((....)))))))......)-------).))..)))..)))).------------- ( -34.40, z-score = -1.56, R) >anoGam1.chr2R 29339698 93 - 62725911 ACGGCGAGCCGAAGAUCAAGGCCGACUUCGAUCAGCUGUACGAAGACAUGCAGCAGGCAUUCCGCAACCUAGAAGAUUAAG------------CGGGCUCUAUCU------------- .(((....))).((((..((.(((.(((.((((.((((((........))))))..((.....)).........)))))))------------))).))..))))------------- ( -27.80, z-score = -0.77, R) >apiMel3.Group9 9325426 115 - 10282195 ACGGUGAAGCAAAAAUCAGAGCUGAUUUUGAUCAAUUACAUGAAGAUAUUCAACAGGCAUUUAGAAAUUUAGAAGAUUAAUUU---UCAAACAUUUAUAUAAAUUAAGUUCUCUAUGU ...((((....(((((((....)))))))......)))).((((....))))....((((..((((.....(((((....)))---)).....((((.......))))))))..)))) ( -13.30, z-score = 1.13, R) >triCas2.ChLG4 10027240 117 + 13894384 ACGGCGAGGCGAAAAUCAAAGCCGAUUUUGACCAGUUGUACGAGGAUAUUCAACAAGCCUUCAGGAACUUGGAGGAUUAAAUUAGUUUUAUUGUAGGCCGUAUAUUUAACCUCACUA- (((((..((..((((((......))))))..)).)))))..((((............(((((((....))))))).((((((..((((......)))).....))))))))))....- ( -25.40, z-score = -0.38, R) >consensus AUGGUGAGGCCAAGAUCAAGGCUGACUUCGAGCAGCUGCACGAGGACCUGCAGCAGGCCUUCAGGAAUCUGGAGGACUAGAG_______CGCGUUGGCCCUACUU_____________ ..(((..(((..........)))...........((((((........)))))).(((((((((....))))))).))..................)))................... (-19.98 = -19.40 + -0.58)

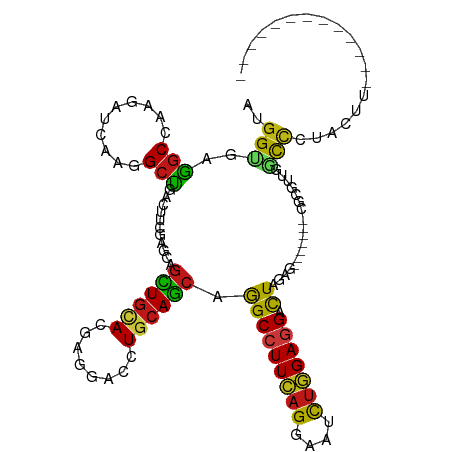

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:36:32 2011