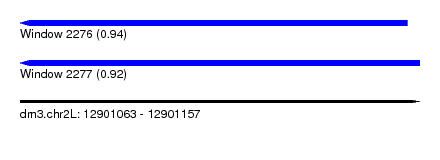
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,901,063 – 12,901,157 |
| Length | 94 |
| Max. P | 0.938840 |

| Location | 12,901,063 – 12,901,154 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 95.97 |
| Shannon entropy | 0.07440 |
| G+C content | 0.28388 |
| Mean single sequence MFE | -21.55 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.12 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.938840 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 12901063 91 - 23011544 GGGAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ...(((...(((((((((.(((..(((....((((((((((((....)))))))))))).....)))...)))..)))))))))...))). ( -20.90, z-score = -2.20, R) >droSim1.chr2L 12693789 91 - 22036055 GGCAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUAACCGUUACCGUUAUUGUUAUUGUUA ...(((...(((((((((.(((.(((.....((((((((((((....)))))))))))).....)))...)))..)))))))))...))). ( -20.80, z-score = -2.00, R) >droSec1.super_16 1061507 91 - 1878335 GGGAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUAACCGUUACCGUUAUUGUUAUUGUUA ...(((...(((((((((.(((.(((.....((((((((((((....)))))))))))).....)))...)))..)))))))))...))). ( -20.90, z-score = -2.16, R) >droAna3.scaffold_12916 3649692 91 + 16180835 GGAGGUGGAGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUAUUGUUACCGUUACCGUUAUUGUUAUUGUUA ((..((((..((((((((((((((...)))))))(((((((((....)))))))))..)))))))..))))..))................ ( -21.70, z-score = -2.85, R) >dp4.chr4_group4 557597 91 - 6586962 GGGAAUGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ((.(((((..(((..((((((...)))))).((((((((((((....))))))))))))...)))..))))).))................ ( -22.50, z-score = -2.70, R) >droPer1.super_46 92778 91 + 590583 GGGAAUGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ((.(((((..(((..((((((...)))))).((((((((((((....))))))))))))...)))..))))).))................ ( -22.50, z-score = -2.70, R) >consensus GGGAACGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ...(((...(((((((((.(((.........((((((((((((....))))))))))))...........)))..)))))))))...))). (-18.64 = -18.12 + -0.53)

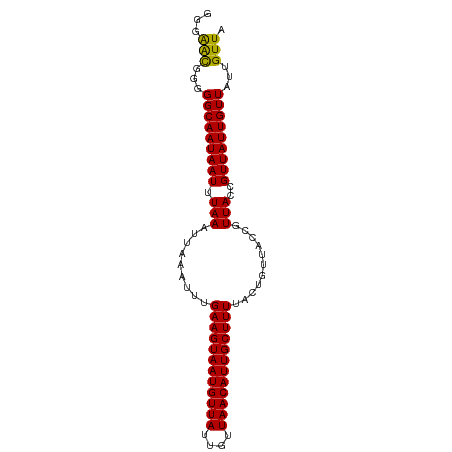
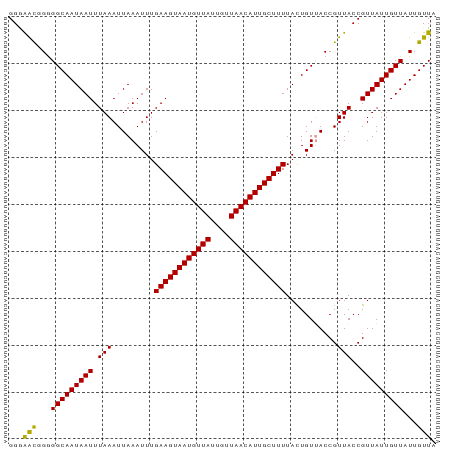
| Location | 12,901,063 – 12,901,157 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 93.76 |
| Shannon entropy | 0.11859 |
| G+C content | 0.29224 |
| Mean single sequence MFE | -22.02 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.12 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.923036 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 12901063 94 - 23011544 GUUGGGAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ......(((...(((((((((.(((..(((....((((((((((((....)))))))))))).....)))...)))..)))))))))...))). ( -20.90, z-score = -1.89, R) >droSim1.chr2L 12693789 94 - 22036055 GUUGGCAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUAACCGUUACCGUUAUUGUUAUUGUUA ..((((((....(((((((((.(((.(((.....((((((((((((....)))))))))))).....)))...)))..))))))))).)))))) ( -21.20, z-score = -1.80, R) >droSec1.super_16 1061507 94 - 1878335 GUUGGGAACUGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUAACCGUUACCGUUAUUGUUAUUGUUA ......(((...(((((((((.(((.(((.....((((((((((((....)))))))))))).....)))...)))..)))))))))...))). ( -20.90, z-score = -1.83, R) >droAna3.scaffold_12916 3649692 91 + 16180835 ---GGAGGUGGAGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUAUUGUUACCGUUACCGUUAUUGUUAUUGUUA ---((..((((..((((((((((((((...)))))))(((((((((....)))))))))..)))))))..))))..))................ ( -21.70, z-score = -2.85, R) >dp4.chr4_group4 557597 94 - 6586962 CGGGGGAAUGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA (((...(((((..(((..((((((...)))))).((((((((((((....))))))))))))...)))..))))).)))............... ( -23.70, z-score = -2.68, R) >droPer1.super_46 92778 94 + 590583 CGGGGGAAUGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA (((...(((((..(((..((((((...)))))).((((((((((((....))))))))))))...)))..))))).)))............... ( -23.70, z-score = -2.68, R) >consensus GUUGGGAACGGGGGCAAUAAUUUAAAUUAAAUUUGAAGUAAUGUUAUUGUUAACAUUGCUUUUACUGUUACCGUUACCGUUAUUGUUAUUGUUA ......(((...(((((((((.(((.........((((((((((((....))))))))))))...........)))..)))))))))...))). (-18.64 = -18.12 + -0.53)
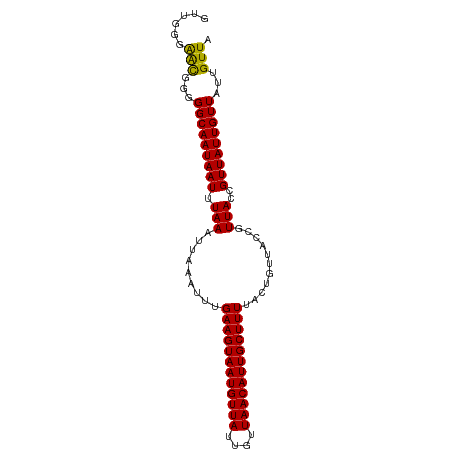


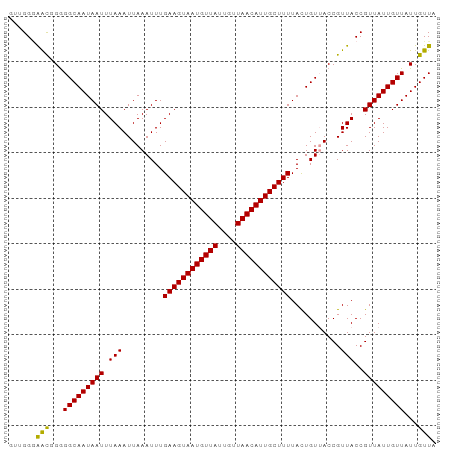
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:36:17 2011