
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,769,045 – 12,769,179 |
| Length | 134 |
| Max. P | 0.750403 |

| Location | 12,769,045 – 12,769,179 |
|---|---|
| Length | 134 |
| Sequences | 9 |
| Columns | 143 |
| Reading direction | forward |
| Mean pairwise identity | 72.92 |
| Shannon entropy | 0.54705 |
| G+C content | 0.43189 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -13.35 |
| Energy contribution | -13.47 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.750403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 12769045 134 + 23011544 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-GGCAGUCCACACAUACAUACGU----AUGUAUAUCCUGCAAUCCACAUC .........((-(((....(((((.(((.((((((((.........-))).))))).))).....((((--((((((...........)))))-))........((((((....))----))))......))))))))))))) ( -30.30, z-score = -1.96, R) >droAna3.scaffold_12916 3537739 119 - 16180835 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-AGGACUUCCCCAGCAC---------------CCUUCCAGCACCCA---- (((((.((((.-.(((((((((...(((.((((((((.........-))).))))).))).)).(((..--..))).....)))))))...))-)))).......)))..---------------..............---- ( -22.51, z-score = -1.19, R) >droEre2.scaffold_4929 13980374 126 - 26641161 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-GGGAGUCCACACAUAC------------GUGUAUCCUGGCAUCCCCAUA .........((-(((((..(((((((((.((((((((.........-))).))))).)))........(--((((((...........)))))-))))))))((((....------------))))....)))))))...... ( -34.30, z-score = -2.64, R) >droYak2.chr2L 9199428 134 + 22324452 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-GGGAGUCUACGCAUACAUACAU----ACAUGCAUAUAGCCAUCCCCAUC .........((-(((((.((((((((((.((((((((.........-))).))))).)))........(--((((((...........)))))-))))))))).((((........----..))))...))).))))...... ( -29.00, z-score = -1.40, R) >droSec1.super_16 943815 138 + 1878335 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-GGGAGUACACACAUGCAUACGUGUGUAUGUAUAUCCUGCAAUCCCCAUC .........((-((((((((((...(((.((((((((.........-))).))))).))).)).(((..--..))).....)))))))..(((-(((.(((((.((((((....)))))).))))).)))))).)))...... ( -37.50, z-score = -2.32, R) >droSim1.chr2L 12567557 138 + 22036055 GCUUCACUUGA-UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAGCA-GGGAGUCCACACAUGCAUACGUAUGUAUGUAUAUCCUGCAAUCCCCAUC ((.......((-(((...((((((((((.((((((((.........-))).))))).)))........(--((((((...........)))))-))))))))).((((((((....)))))))))))))..)).......... ( -35.40, z-score = -2.21, R) >droWil1.scaffold_180708 6790475 126 - 12563649 UCUUCUCUAUGAUGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACAACAUUAACUAUCAUAGU-AUGUGUGUGUGUGUUUGUGUGCUCUUCCACUC------------- ..................((((...((....))...((.(((((((-(((((((((((((....(((..--..)))....))))).((((...))))-.....)))))))))))))).).)).))))...------------- ( -29.70, z-score = -1.47, R) >droMoj3.scaffold_6500 25776903 116 + 32352404 GCUGCACUUCA-UUUUAAUGGAUUCGCACUCACAAUGAACCGCAAG-CAUAUGCAAAUGUCUCGAGCAC--CUGCUAACGACAUUAACUAGCAUAGUGGAGUGUGG-UGUGAGAGUGGGGU---------------------- (((.(((((((-(....)))))(((((((.((((.....((((..(-(....)).((((((...(((..--..)))...))))))..........))))..)))))-)))))).))).)))---------------------- ( -36.80, z-score = -1.57, R) >droGri2.scaffold_15252 9826117 112 + 17193109 GCCGCACUUCAGUUUUAAUGUAUUCGCACUCGCACUGAACCGCAAAACAGAUGCAAAUGUCUCGAGCAUUACAGCUAACAGCAUUAACUAGCAUAGUGAAGUGUAU-AAUGUC------------------------------ ...(((((((((((.((((((..(((.((..((((((..........))).)))....))..))))))))).))))....((........)).....)))))))..-......------------------------------ ( -23.50, z-score = 0.05, R) >consensus GCUUCACUUGA_UGUUAAUGGAUUCGCACUUGCAAUGAACCACAAA_CAUAUGCAAAUGUCUCGAGCAC__CUGCUAACAACAUUAACUAGCA_GGGAGUCCACACAUACAUACGU____A_GU_UAUCCUGC_AUCCCCAU_ ......((.....(((((((.....(((.((((((((..........))).))))).)))....(((......))).....)))))))......))............................................... (-13.35 = -13.47 + 0.11)

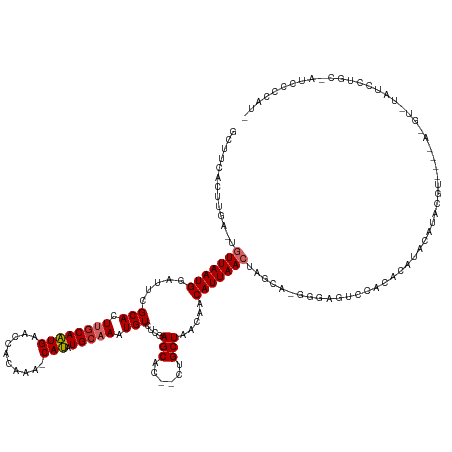

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:35:48 2011