
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,457,225 – 12,457,316 |
| Length | 91 |
| Max. P | 0.693955 |

| Location | 12,457,225 – 12,457,316 |
|---|---|
| Length | 91 |
| Sequences | 9 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 67.54 |
| Shannon entropy | 0.61798 |
| G+C content | 0.51517 |
| Mean single sequence MFE | -28.46 |
| Consensus MFE | -15.03 |
| Energy contribution | -14.85 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.36 |
| Mean z-score | -0.77 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.693955 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 12457225 91 + 23011544 ------GGCGCAGCAGCGCGAUAUUUAAAAGAACU-----CCUAUACUCGCUUCGAGU-----UAACAGCGGACGUUAACAGCGC-CAGCCGCAGCCGCUGUUAGCGU ------.((((....))))..............((-----.((..(((((...)))))-----....)).))((((((((((((.-..((....)))))))))))))) ( -28.80, z-score = -0.45, R) >droSim1.chr2L 12263655 91 + 22036055 ------GGCGCAGCAGCGCGAUAUUUAAAAGAACU-----UCUAUACUCGCUUCGAGU-----UAACAGCGGACGUUAACAGCGC-CAGCCGCAGCCGCUGUUAGCGU ------.((((....))))..............((-----.((..(((((...)))))-----....)).))((((((((((((.-..((....)))))))))))))) ( -28.80, z-score = -0.38, R) >droSec1.super_16 648089 91 + 1878335 ------GGCGCAGCAGCGCGAUAUUUAAAAGAACU-----UCUAUACUCGCUUCGAGU-----UAACAGCGGACGUUAACAGCGC-CAGCCGCAGCCGCUGUUAGCGU ------.((((....))))..............((-----.((..(((((...)))))-----....)).))((((((((((((.-..((....)))))))))))))) ( -28.80, z-score = -0.38, R) >droYak2.chr2L 8894142 91 + 22324452 ------GGCGCAGCAGCGCACCUUUUAAAAGAGCU-----CCUAUACUCGCUUCGAGU-----UAACAGCGGACGUUAACAGCGC-CAGCCGCAGCCGCUGUUAGCGU ------.((((....)))).............(((-----.....(((((...)))))-----....)))..((((((((((((.-..((....)))))))))))))) ( -31.60, z-score = -1.12, R) >droEre2.scaffold_4929 13679166 91 - 26641161 ------GGCGCAGCAGCGCGAUAUUUAAAAGAGCU-----CCUAUACUCGCUUCGAGU-----UAACAGCGGACGUUAACAGCGC-CAGCCGCAGCCGCUGUUAGCGU ------.((((....)))).............(((-----.....(((((...)))))-----....)))..((((((((((((.-..((....)))))))))))))) ( -31.60, z-score = -0.86, R) >dp4.chr4_group3 3146931 108 + 11692001 UGUUUCCGCGCAGCAGCGCCUCUUUAAAACUAACUGUAAUUCUUCAUCUUCUUCUGUUAAGCAUAACAGCAGACGUUAACAGCAUGCAGCCGCAGCCAUUGUUAACGU .......((((....))))..............................(((.((((((....)))))).)))((((((((((.(((....)))))...)))))))). ( -25.60, z-score = -1.36, R) >droPer1.super_1 4629655 108 + 10282868 UGUUUCCGCGCAGCAGCGCCUCUUUAAAACUAACUGUAAUUCUUCAUCUUCUUCUGUUAAGCAUAACAGCAGACGUUAACAGCAUGCAGCCGCAGCCAUUGUUAACGU .......((((....))))..............................(((.((((((....)))))).)))((((((((((.(((....)))))...)))))))). ( -25.60, z-score = -1.36, R) >droVir3.scaffold_13246 1654655 94 + 2672878 ----UUGGCGCAGCAGCGCACUGUUAAAUAAU-AA-----UAAAAACACUCUGUUAAAGCGUUCGCUAACAGCUGUUGGCCAGCCGCAGCCA--G--GCUGUUAACGC ----..(((.(((((((....(((((.....)-))-----)).........((((..(((....)))))))))))))))))((((.......--)--)))........ ( -27.00, z-score = 0.16, R) >droMoj3.scaffold_6500 28083406 92 - 32352404 ----UUGGCGCAGCAGCGCUCUGUUAAAUAAAGAA-----UGAAAACACUCUGCGAAAGCCGACGUUAACA---GUUGGCCAGCCGCAGCCAUAG--GCUGUUAAC-- ----..((((((((((....)))))......(((.-----((....)).)))))....((((((.......---))))))..)))((((((...)--)))))....-- ( -28.30, z-score = -1.17, R) >consensus ______GGCGCAGCAGCGCGAUAUUUAAAAGAACU_____UCUAUACUCGCUUCGAGU_____UAACAGCGGACGUUAACAGCGC_CAGCCGCAGCCGCUGUUAGCGU .......((((....)))).....................................................((((((((((................)))))))))) (-15.03 = -14.85 + -0.19)


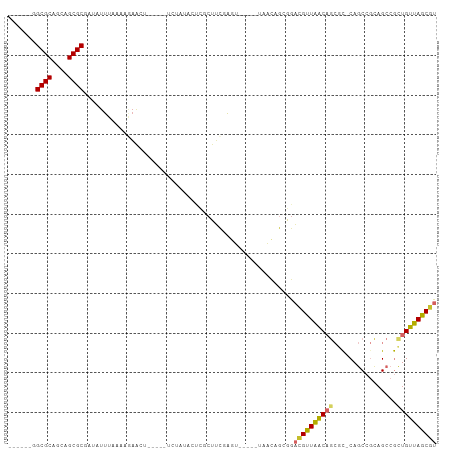
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:35:13 2011