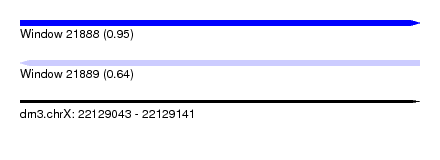

| Sequence ID | dm3.chrX |
|---|---|
| Location | 22,129,043 – 22,129,141 |
| Length | 98 |
| Max. P | 0.945932 |

| Location | 22,129,043 – 22,129,141 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 52.83 |
| Shannon entropy | 0.76604 |
| G+C content | 0.44824 |
| Mean single sequence MFE | -28.32 |
| Consensus MFE | -9.02 |
| Energy contribution | -10.77 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.945932 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
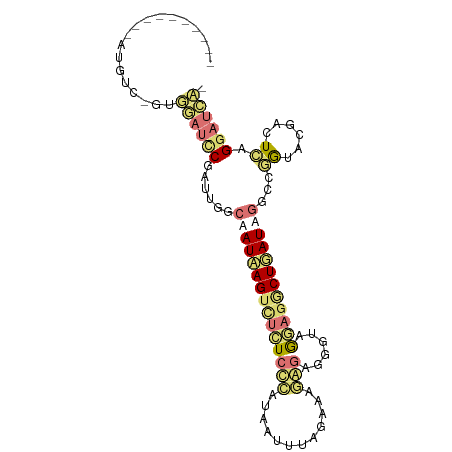
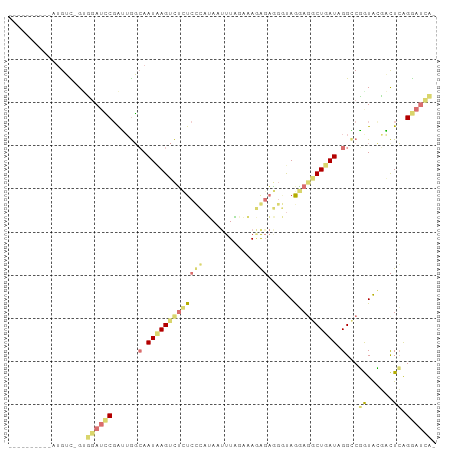
>dm3.chrX 22129043 98 + 22422827 UUGUCUUAUAAUGUC-GUUGAUCCGAUUGGCAAUAAGUCUCUCCCAUAAUUUAGAAAGGGAGGGUAGGAGGCUUAUAGGACGAUACGACUCAGGAUCAA ..(((((((((.(((-((..((...))..)).......(((((((............))))))).....))))))))))))(((.(......).))).. ( -26.90, z-score = -1.17, R) >droSim1.chrU 15302561 98 + 15797150 UUGUCUUAUAAUGUC-GUUGAUCCGAUUGGCAAUAAGUCUCUCCCAUAAUUUAGAAAGGGAGGGUAAGAGGCUUAUAGGCCGGUACGACUCAGGAUCAU ...............-..((((((.((((((.(((((((((((((................)))..))))))))))..))))))........)))))). ( -29.09, z-score = -1.76, R) >anoGam1.chrU 11629019 89 + 59568033 ----------AUGUUUGCGAAUUCUGAUAGCAAUCAGUUUUUCCAAAAGUUUUACCAAACAACUUAGAAGACUGAUUGGUCUGUAACUUUCAGGAUUCG ----------.......((((((((((.((((((((((((((....(((((.........))))).))))))))))))........)).)))))))))) ( -24.40, z-score = -3.30, R) >triCas2.chrUn_101 67239 87 - 68148 ----------GAGGUGGAGGAGGCGGGAGCGAAUGAGACGUUCUGCGACCUUUCCCACAGCCGACAGCAAGCUGAUUGGCCCGUGGUUCUGGGAAAA-- ----------...((((..((((((((((((.......)))))).)).))))..)))).((((((((....))).)))))(((......))).....-- ( -32.90, z-score = -1.19, R) >consensus __________AUGUC_GUGGAUCCGAUUGGCAAUAAGUCUCUCCCAUAAUUUAGAAAGAGAGGGUAGGAGGCUGAUAGGCCGGUACGACUCAGGAUCA_ ..................((((((......(((((((((((((((............)))......))))))))))))...((......)).)))))). ( -9.02 = -10.77 + 1.75)

| Location | 22,129,043 – 22,129,141 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 52.83 |
| Shannon entropy | 0.76604 |
| G+C content | 0.44824 |
| Mean single sequence MFE | -22.18 |
| Consensus MFE | -6.57 |
| Energy contribution | -7.45 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.57 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.30 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.644750 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
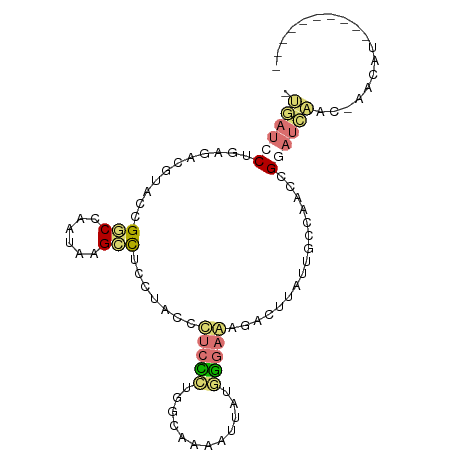
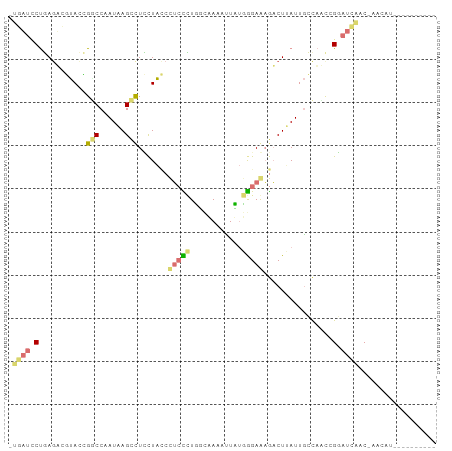
>dm3.chrX 22129043 98 - 22422827 UUGAUCCUGAGUCGUAUCGUCCUAUAAGCCUCCUACCCUCCCUUUCUAAAUUAUGGGAGAGACUUAUUGCCAAUCGGAUCAAC-GACAUUAUAAGACAA ((((((((((((((((......)))............(((((............))))).)))))).........))))))).-............... ( -19.10, z-score = -0.68, R) >droSim1.chrU 15302561 98 - 15797150 AUGAUCCUGAGUCGUACCGGCCUAUAAGCCUCUUACCCUCCCUUUCUAAAUUAUGGGAGAGACUUAUUGCCAAUCGGAUCAAC-GACAUUAUAAGACAA .(((((((((((((((..(((......)))...))).(((((............))))).)))))).........))))))..-............... ( -24.60, z-score = -2.32, R) >anoGam1.chrU 11629019 89 - 59568033 CGAAUCCUGAAAGUUACAGACCAAUCAGUCUUCUAAGUUGUUUGGUAAAACUUUUGGAAAAACUGAUUGCUAUCAGAAUUCGCAAACAU---------- (((((.((((.((........((((((((.(((((((..((((....)))).)))))))..)))))))))).)))).))))).......---------- ( -23.20, z-score = -3.19, R) >triCas2.chrUn_101 67239 87 + 68148 --UUUUCCCAGAACCACGGGCCAAUCAGCUUGCUGUCGGCUGUGGGAAAGGUCGCAGAACGUCUCAUUCGCUCCCGCCUCCUCCACCUC---------- --.(((((((.(.(....)(((...(((....)))..)))).)))))))((..((.((.((.......)).))..))..))........---------- ( -21.80, z-score = 0.59, R) >consensus _UGAUCCUGAGACGUACCGGCCAAUAAGCCUCCUACCCUCCCUGGCAAAAUUAUGGGAAAGACUUAUUGCCAACCGGAUCAAC_AACAU__________ .((((((...........(((......))).......(((((............)))))................)))))).................. ( -6.57 = -7.45 + 0.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:07:43 2011