
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,301,943 – 12,302,108 |
| Length | 165 |
| Max. P | 0.527757 |

| Location | 12,301,943 – 12,302,108 |
|---|---|
| Length | 165 |
| Sequences | 6 |
| Columns | 185 |
| Reading direction | reverse |
| Mean pairwise identity | 67.91 |
| Shannon entropy | 0.54934 |
| G+C content | 0.54379 |
| Mean single sequence MFE | -56.99 |
| Consensus MFE | -22.23 |
| Energy contribution | -22.82 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.527757 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 12301943 165 - 23011544 AAAGAAGGCAGGAUUCACAGAUA-GGCGCAUCCCUGCUU----GCUGUCCUCGCUGUCAGCUGCACUUUGAGCUGAAUUUCUUUUUC----AUUCGAAGUGAAAGGAUGUAACAGACGGAGGACA---UCUCCUUCGAGUCC--------GAAUCGUCGGACUCCAUCCGCUCGCUCCCGGGGCA ..((.(((((((((.(.(.....-.).).)).)))))))----.))(((((.(..((.(((..(((((((((.((((.......)))----))))))))))...(((((........((((((..---..))))))((((((--------((....)))))))))))))))).))..).))))). ( -62.80, z-score = -2.38, R) >droSim1.chr2L 12102379 142 - 22036055 -------------------AAUA-GGCGCAUCCCUGCUC--------UCCUCACUGUCAGCUGCACUUUGAGCUGAAUUUCUUUUUC----AUUCGCAGGGGAAGGAUGUAACAGACGGAGGACA---UCUCCUUCGAGUCCGA----AUGAAUCGUCGGACUCCAUC----CGCUGCCGGGGCA -------------------....-(((((.(((((((..--------.........((((((........))))))...........----....)))))))..(((((........((((((..---..))))))((((((((----........))))))))))))----))).)))...... ( -53.67, z-score = -2.37, R) >droSec1.super_16 491634 143 - 1878335 -------------------GAUAGGGCGCAUCCCUGCUC--------UCCUCACUGUCAGCUGCACUUUGAGCUGAAUUUCUUUUUC----AUUCGCAGGGGAAGGAUGUAACAGACGGAGGACA---UCUCCUUCGAGUCCGA----AUGAAUCGUCGGACUCCAUC----CGCUGCCGGGGCA -------------------.....(((((.(((((((..--------.........((((((........))))))...........----....)))))))..(((((........((((((..---..))))))((((((((----........))))))))))))----))).)))...... ( -53.47, z-score = -1.73, R) >droYak2.chr2L 8732435 166 - 22324452 ----CAGAGAGGAUUCACAGAUA-GGCGCAUCCUUGCUGUCCAGCUGUCCUCGCUGUCAGCUGCACUUUGAGCUGAAUUUCUUUUUC----GUUCGAAGGGAAAGGAUGUAACAGACGGAGGACA---UCUCCUUCGAGUCCGG---UCCGAAUCGUCGGACUCCAUC----CGCUCCUGGGGCA ----(((.(((((....(((...-((((((....))).)))...)))))))).)))...(((.((((((((((.(((.......)))----))))))))(((..(((((........((((((..---..))))))((((((((---.(......)))))))))))))----)..))))).))). ( -64.00, z-score = -1.66, R) >droEre2.scaffold_4929 13524848 147 + 26641161 ----CAGCAAGGAUUCGCAGAUA-GGCGCAUCCUUGCUGUCCAGCUGUCCUCGCUGCCAGCUGCACUUUGAGCAGAAUUUCUUUUUC----ACUCCAAGGCGCAGGAUGUAACAGACGGAGGACA---UCUCCUUCGAGUCCGA----AUCCGCUCCCGGGCA---------------------- ----.(((((((((.(((.....-.))).)))))))))((((((.(((((((.(((.((.((((.(((((((..(((......))).----.)).))))).))))..))...)))...)))))))---.)).....((((....----....))))..)))).---------------------- ( -54.30, z-score = -1.80, R) >droAna3.scaffold_12916 3084775 155 + 16180835 AGGAAAGGAAGGACGAAAAGAUA-GGCGCAUCCUUGCUG--------UCCCCACUGUCAGCUGCACUUUGAAGCGAGUUUCCUUUUCGCCAAUUCCGGGUGGUGGAAGGAUGGUGACGGGGAACAAGGACAUCUCCGAAUCGGAGUCCAUCUGCUCCCGGGCCA--------------------- .((((((((((..((.......(-((.(((.....((((--------.(......).))))))).))).....))..))))))))))......((((((.((..((.((((..(((((((((.........))))))..)))..)))).))..))))))))...--------------------- ( -53.70, z-score = 0.41, R) >consensus _____AG__AGGAUUCACAGAUA_GGCGCAUCCCUGCUG________UCCUCACUGUCAGCUGCACUUUGAGCUGAAUUUCUUUUUC____AUUCGAAGGGGAAGGAUGUAACAGACGGAGGACA___UCUCCUUCGAGUCCGA____AUGAAUCGUCGGACUCCAUC____CGCU_CCGGGGCA ........................(..((((((.(((.........................)))(((((((..(((.......))).....))))))).....))))))..)...(((((((.......))))))).(((((..............)))))....................... (-22.23 = -22.82 + 0.59)

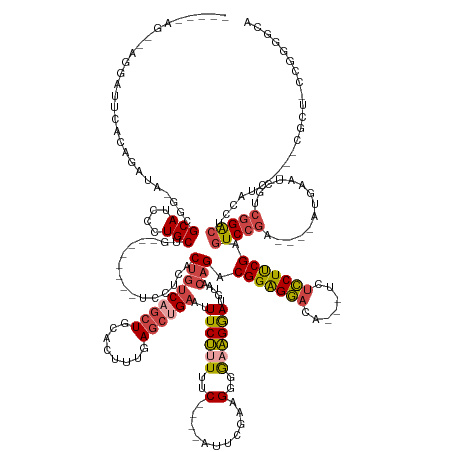

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:34:51 2011