| Sequence ID | dm3.chrX |
|---|---|
| Location | 22,045,013 – 22,045,120 |
| Length | 107 |
| Max. P | 0.624847 |

| Location | 22,045,013 – 22,045,120 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 59.94 |
| Shannon entropy | 0.56756 |
| G+C content | 0.46080 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -14.11 |
| Energy contribution | -14.90 |
| Covariance contribution | 0.79 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.624847 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 22045013 107 + 22422827 GCUCAGCUAACAGAGAACAAAGUGCAUUUUACGAAUCUGCCCUUAGGGCUAGAG-CUGAUCUCUCGCUCGCU--CUCAUUUGCUGGCCUGAGUUAAGUAGGAAGAAUACC (((((((((.((((......((.((.............((((...))))..(((-(.((....)))))))).--))..)))).))).))))))................. ( -26.00, z-score = 0.91, R) >droSim1.chr3h_random 1016533 93 + 1452968 ------UAAACGACUAAGAAAACUUAGUGUACGACCCUGCCCAUUAGGCUGGUGUCUUGUCUCUCACUUGUUAACGCAUAGGCCGGCGAGAGAUAAGAA----------- ------...((.((((((....))))))))...(((..(((.....))).))).(((((((((((.((.(((........))).)).))))))))))).----------- ( -31.00, z-score = -3.29, R) >droYak2.chrU 12127624 106 - 28119190 ACUCAGAUAACAGAGUACAAAGCGCACUUUACUUCUCUGCCCUAGGGGCUAGAG-CUUACCUCACGCUCGCU--CUUAUU-GCUGGCUUGAGUUAAGUAAUAAGAACAAU ((((........))))....((((..........((((((((...)))).))))-.........)))).(.(--((((((-((((((....))).)))))))))).)... ( -27.61, z-score = -0.79, R) >consensus _CUCAGAUAACAGAGAACAAAGCGCACUUUACGACUCUGCCCUUAGGGCUAGAG_CUUAUCUCUCGCUCGCU__CUCAUU_GCUGGCCUGAGUUAAGUA__AAGAA_A__ ..................................(((((((.....))).)))).((((.((((.(((.((..........)).))).)))).))))............. (-14.11 = -14.90 + 0.79)

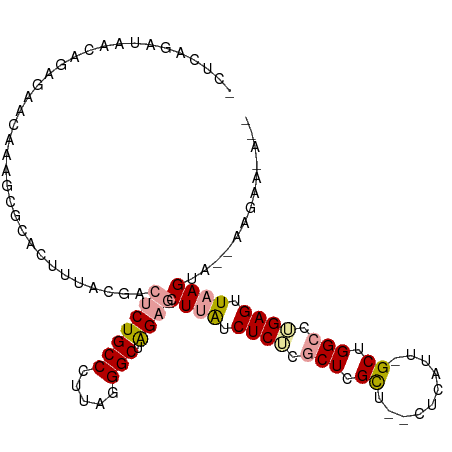
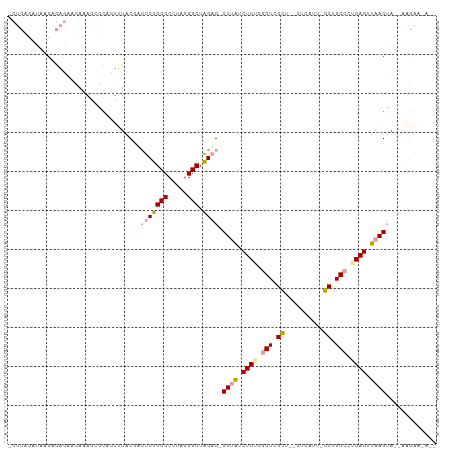
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:07:32 2011