
| Sequence ID | dm3.chrX |
|---|---|
| Location | 21,855,442 – 21,855,541 |
| Length | 99 |
| Max. P | 0.660232 |

| Location | 21,855,442 – 21,855,541 |
|---|---|
| Length | 99 |
| Sequences | 9 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 79.19 |
| Shannon entropy | 0.37705 |
| G+C content | 0.52628 |
| Mean single sequence MFE | -26.25 |
| Consensus MFE | -20.01 |
| Energy contribution | -20.06 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.12 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.660232 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
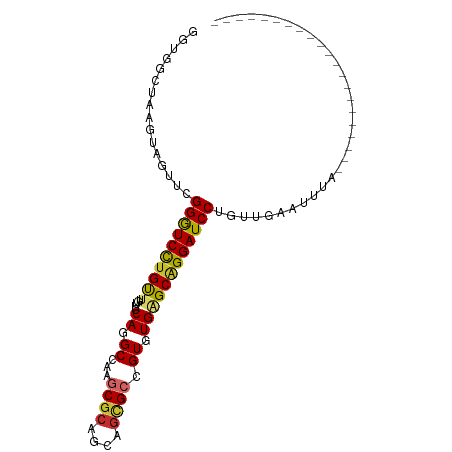
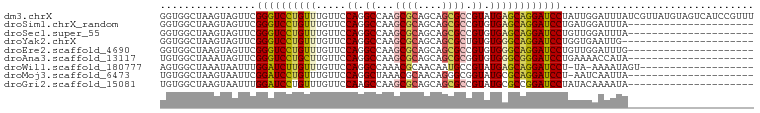
>dm3.chrX 21855442 99 + 22422827 GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCCGUAUGAGCAGGAUCCUAUUGGAUUUAUCGUUAUGUAGUCAUCCGUUU ((((((((.((((...((((((((((..(....(((...((......)))))..)..)))))))))).....((....)).)))).))))))))..... ( -34.30, z-score = -2.68, R) >droSim1.chrX_random 5658279 78 + 5698898 GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCCGUGUGAGCAGGAUCCUGAUGGAUUUA--------------------- ......(((((...((((((((((((..(....(((...((......)))))..)..))))))).)))))...)))))--------------------- ( -27.10, z-score = -0.87, R) >droSec1.super_55 11441 78 + 187931 GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCCGUGUGAGCAGGAUCCUGUUGGAUUUA--------------------- ......(((((....(((((((((((..(....(((...((......)))))..)..))))))).))))....)))))--------------------- ( -25.10, z-score = -0.27, R) >droYak2.chrX 21300977 77 + 21770863 GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCUGUGUGGGCAGGAUCCUGGUGAAUUG---------------------- ..........(((((((((((((((.....(((.((..(((((....))))))).))))))))))))....))))))---------------------- ( -28.70, z-score = -1.11, R) >droEre2.scaffold_4690 18233779 78 + 18748788 GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCCGUGUGGGCAGGAUCCUGUUGGAUUUG--------------------- ................(((((((((.....(((.((...((((....)))).)).))))))))))))...........--------------------- ( -26.30, z-score = -0.20, R) >droAna3.scaffold_13117 3829493 78 - 5790199 UGUGGCUAAAUAGUUCGGGUCCUGCUUGUUCCAGGCCAAGCGCAGCAGCGCGGUGUGGGCGGGAUCCUGAAAACCAUA--------------------- (((((........((((((((((((.....(((.(((..((((....))))))).))))))))).))))))..)))))--------------------- ( -33.20, z-score = -2.09, R) >droWil1.scaffold_180777 2456024 78 + 4753960 AGUGGCUAAAUAAUUUGGAUCUUGUUUGUUCCAGGCCAAACGCAACAAUGCCGUAUGAGCAGGAUCCU-UA-AAAAUAGU------------------- ....((((........((((((((((..(....(((.............)))..)..)))))))))).-..-....))))------------------- ( -20.78, z-score = -1.95, R) >droMoj3.scaffold_6473 3338748 77 + 16943266 UGUGGCUAAGUAAUUCGGAUCCUGUUUGUUCCAGGCUAAACGCAACAGGGCGGUAUGCGCAGGAUCCU-AAUCAAUUA--------------------- ................(((((((((.........((....(((......)))....))))))))))).-.........--------------------- ( -20.10, z-score = -0.53, R) >droGri2.scaffold_15081 3105341 78 - 4274704 UGUGGCUAAGUAAUUUGGAUCCUGUUUGUUCCAAGCCAAGCGCAGCAGCGCCGUAUGCGCCGGAUCCUAUACAAAAUA--------------------- .........(((....((((((.(((((...)))))...(((((((......)).))))).))))))..)))......--------------------- ( -20.70, z-score = -0.42, R) >consensus GGUGGCUAAGUAGUUCGGGUCCUGUUUGUUCCAGGCCAAGCGCAGCAGCGCCGUGUGAGCAGGAUCCUGUUGAAUUUA_____________________ ................((((((((((.....((.((...((((....)))).)).))))))))))))................................ (-20.01 = -20.06 + 0.04)


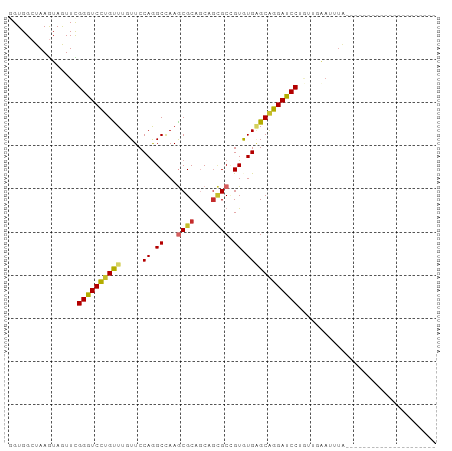
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:07:24 2011