
| Sequence ID | dm3.chrX |
|---|---|
| Location | 21,668,095 – 21,668,189 |
| Length | 94 |
| Max. P | 0.784075 |

| Location | 21,668,095 – 21,668,189 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 77.22 |
| Shannon entropy | 0.31703 |
| G+C content | 0.53244 |
| Mean single sequence MFE | -24.97 |
| Consensus MFE | -14.57 |
| Energy contribution | -14.90 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.784075 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 21668095 94 - 22422827 AGCGAGAGUUACCGGUUCCAACCGGCACACAGACCCUCCAGCGAAGCCACCCCGGCUUACUCUGGAGUUGGUACAAAGUAACCAAUGUAUAAAC (((....))).(((((....)))))...(((....((((((.((((((.....))))).).))))))(((((........))))))))...... ( -30.30, z-score = -2.96, R) >droSim1.chrU 13095512 93 + 15797150 -GCGUGAGUUACCGGUGCCAACCGGCACACAGACCCUCCAGCAAAGCCACCCCGGCUUACCCUGGAAUUGGUACAAAGUAACCAAAACAUAAAC -...((.(((((..(((((....)))))....(((.(((((..(((((.....)))))...)))))...))).....))))))).......... ( -28.90, z-score = -3.06, R) >droSec1.super_49 61646 72 + 245908 -GCGCGAGUUACCGGUUCCAACCG---------------------GCCACCCCGUCUUACCCUGGAAUUGGUACAAAGUAACCAAAACCUAAAC -(((.(.((..(((((....))))---------------------)..)).))))........((..(((((........)))))..))..... ( -15.70, z-score = -0.59, R) >consensus _GCGAGAGUUACCGGUUCCAACCGGCACACAGACCCUCCAGC_AAGCCACCCCGGCUUACCCUGGAAUUGGUACAAAGUAACCAAAACAUAAAC .....(.(((((((((....)))).....................((((..((((......))))...)))).....))))))........... (-14.57 = -14.90 + 0.33)


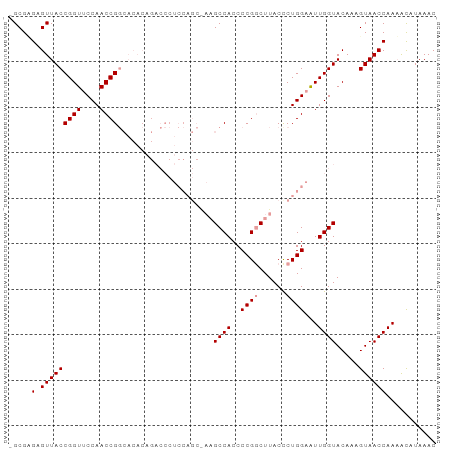
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:07:14 2011