
| Sequence ID | dm3.chrX |
|---|---|
| Location | 21,150,767 – 21,150,860 |
| Length | 93 |
| Max. P | 0.970494 |

| Location | 21,150,767 – 21,150,859 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 75.72 |
| Shannon entropy | 0.39743 |
| G+C content | 0.45003 |
| Mean single sequence MFE | -20.00 |
| Consensus MFE | -8.43 |
| Energy contribution | -10.37 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.32 |
| Mean z-score | -3.06 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.78 |
| SVM RNA-class probability | 0.967529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
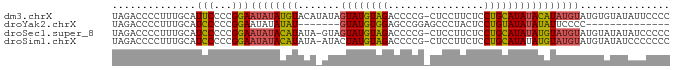
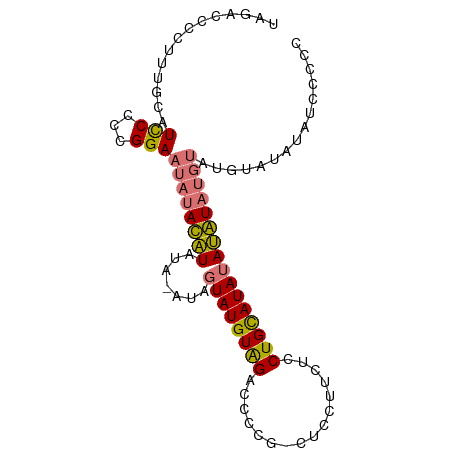
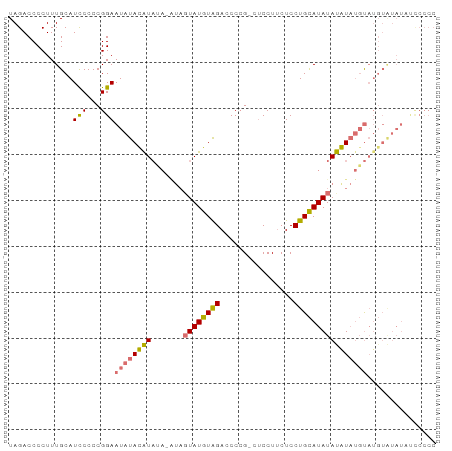
>dm3.chrX 21150767 92 - 22422827 UAGACCCCUUUGCAUUCCCCGGAAUAUAUGUACAUAUAGUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUACAUAUGUAUGUGUAUAUUCCCC ....................(((((((((((((((((.((((((((......-.........))))))))..))))))))..))))))))).. ( -22.96, z-score = -3.87, R) >droYak2.chrX 20326983 72 - 21770863 UAGACCCCUUUGCAUCCCCCGGAAUAUAUAU-------GUAUGUGGAGCCGGAGCCCUACUCCUGUAUAUAUAUUCCCC-------------- ....................(((((((((((-------(((.((((.((....)).))))...))))))))))))))..-------------- ( -17.80, z-score = -2.01, R) >droSec1.super_8 3456560 91 - 3762037 UAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA-GUAGUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUAUGUAUGUAUGUAUAUAUCCCCC ....................((((((((((((((-...((((((((......-.........)))))))).....))))))))))).)))... ( -20.26, z-score = -2.82, R) >droSim1.chrX 16319357 91 - 17042790 UAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA-AUACUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUAUGUAUGUAUGUAUAUCCCCCCC ....................((.(((((((((((-.((((((((((......-.........)))))))...)))))))))))))).)).... ( -18.96, z-score = -3.55, R) >consensus UAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA_AUAGUAUGUAGACCCCG_CUCCUUCUCCUGCAUAUAUAUAUGUAUGUAUAUAUCCCCC ..............(((...)))(((((((((((....((((((((................)))))))).....)))))))))))....... ( -8.43 = -10.37 + 1.94)
| Location | 21,150,767 – 21,150,860 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 75.99 |
| Shannon entropy | 0.39320 |
| G+C content | 0.44481 |
| Mean single sequence MFE | -20.00 |
| Consensus MFE | -8.43 |
| Energy contribution | -10.37 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.32 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.83 |
| SVM RNA-class probability | 0.970494 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
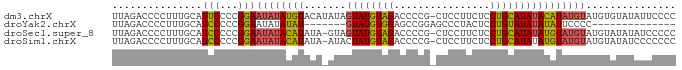
>dm3.chrX 21150767 93 - 22422827 UUAGACCCCUUUGCAUUCCCCGGAAUAUAUGUACAUAUAGUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUACAUAUGUAUGUGUAUAUUCCCC .....................(((((((((((((((((.((((((((......-.........))))))))..))))))))..))))))))).. ( -22.96, z-score = -3.91, R) >droYak2.chrX 20326983 73 - 21770863 UUAGACCCCUUUGCAUCCCCCGGAAUAUAUAU-------GUAUGUGGAGCCGGAGCCCUACUCCUGUAUAUAUAUUCCCC-------------- .....................(((((((((((-------(((.((((.((....)).))))...))))))))))))))..-------------- ( -17.80, z-score = -2.00, R) >droSec1.super_8 3456560 92 - 3762037 UUAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA-GUAGUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUAUGUAUGUAUGUAUAUAUCCCCC .....................((((((((((((((-...((((((((......-.........)))))))).....))))))))))).)))... ( -20.26, z-score = -2.91, R) >droSim1.chrX 16319357 92 - 17042790 UUAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA-AUACUAUGUAGACCCCG-CUCCUUCUCCUGCAUAUAUGUAUGUAUGUAUAUCCCCCCC .....................((.(((((((((((-.((((((((((......-.........)))))))...)))))))))))))).)).... ( -18.96, z-score = -3.63, R) >consensus UUAGACCCCUUUGCAUCCCCCGGAAUAUACAUAUA_AUAGUAUGUAGACCCCG_CUCCUUCUCCUGCAUAUAUAUAUGUAUGUAUAUAUCCCCC ...............(((...)))(((((((((((....((((((((................)))))))).....)))))))))))....... ( -8.43 = -10.37 + 1.94)
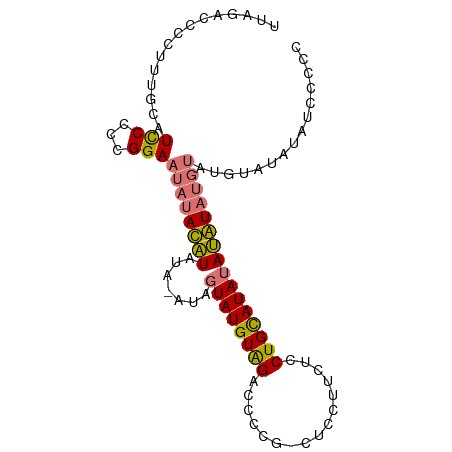
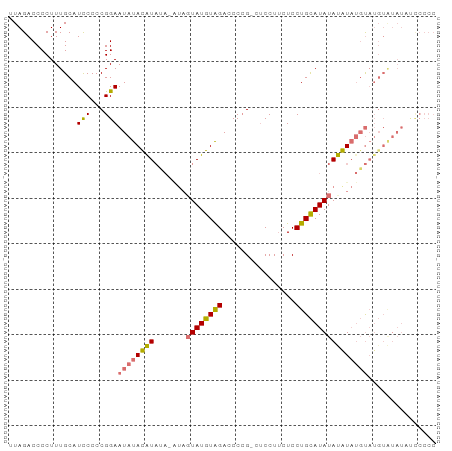
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:06:10 2011