| Sequence ID | dm3.chrX |
|---|---|
| Location | 21,127,494 – 21,127,575 |
| Length | 81 |
| Max. P | 0.956974 |

| Location | 21,127,494 – 21,127,575 |
|---|---|
| Length | 81 |
| Sequences | 11 |
| Columns | 86 |
| Reading direction | reverse |
| Mean pairwise identity | 82.44 |
| Shannon entropy | 0.38424 |
| G+C content | 0.44093 |
| Mean single sequence MFE | -24.58 |
| Consensus MFE | -12.96 |
| Energy contribution | -13.34 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.956974 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 21127494 81 - 22422827 ---ACACAUUUGGCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG--CGCAACUGCCAAAGGUAAUGGCCAAAAAUGGAACA ---..((((((..(((((((.....))))))).))))))...(((((((--((....))))...(((....))).....))))).. ( -25.60, z-score = -2.79, R) >droYak2.chrX 20301353 84 - 21770863 GCCAGACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG--CGCAACUGCCAAAGGUAAUGGGCAAAAACGGGACA (((.(((((((..(((((((.....))))))).)))))))(((.((.((--((....))))...)).))).)))............ ( -27.80, z-score = -2.68, R) >droEre2.scaffold_4690 17657007 84 - 18748788 GCCAGACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG--CGCAACUGCCAAAGGUAAUGGCCAAAAACGGGCCA (((.(((((((..(((((((.....))))))).))))))).......((--((....))))...)))..(((((.......))))) ( -32.90, z-score = -4.10, R) >droSec1.super_8 3433071 81 - 3762037 ---ACACAUUUGGCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUGCAGG--CGCAACUGCCAAAGGUAAUGGCCAAAAACGGAACA ---.....((((((.(((..(((((((((((....))))))))))).((--((....)))).......)))))))))......... ( -28.40, z-score = -3.74, R) >droSim1.chrX_random 5473122 81 - 5698898 ---ACACAUUUGGCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUGCAGG--CGCAACUGGCAAAGGUAAUGGCCAAAACCGGAACA ---.....((((((.(((..(((((((((((....)))))))))))...--.((.....)).......)))))))))......... ( -25.00, z-score = -2.27, R) >droAna3.scaffold_13334 454190 79 + 1562580 -----ACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG--CGCAACUGCCAAAGGUAAUGGCCAAAAACGGGACA -----((((((..(((((((.....))))))).))))))...((((.((--((....))))...(((....)))......)))).. ( -23.90, z-score = -2.17, R) >dp4.chrXL_group1a 44801 77 + 9151740 -----ACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGGGCGAAAACUGCCAAAGGUAAUGGCCAAAAACAG---- -----((((((..(((((((.....))))))).))))))((((.((..((((.....))))...)).))))...........---- ( -22.40, z-score = -2.76, R) >droPer1.super_15 680996 77 + 2181545 -----ACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGGGCGAAAACUGCCAAAGGUAAUGGCCAAAAACAG---- -----((((((..(((((((.....))))))).))))))((((.((..((((.....))))...)).))))...........---- ( -22.40, z-score = -2.76, R) >droWil1.scaffold_180702 117621 80 - 4511350 -----ACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG-CACUAGCUGCCAAAGGUAAUGGACAAAAAGGGCAUA -----.....((((((((((.....)))))).....(((((((.((.((-((.....))))...)).))).)))).....)))).. ( -22.60, z-score = -1.90, R) >droVir3.scaffold_12970 7878153 72 - 11907090 ---ACACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG----CAA-------CGGUCCGGGACACAGAAGGUACG ---..((((((..(((((((.....))))))).))))))...((((.((----(..-------..))).))))............. ( -18.90, z-score = -1.63, R) >droMoj3.scaffold_6482 1075314 77 + 2735782 ---ACACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGACAUUUCCAGG----CAAUGG--ACCAGUUCGGGACACAGAAGGUGGA ---.(((..(((((((((((.....))))))((((......))))..))----)))((.--.((.....))..))......))).. ( -20.50, z-score = -1.32, R) >consensus ___ACACAUUUGCCACAUCAGCAAAUGAUGUGAAAAUGUCAUUUCCAGG__CGCAACUGCCAAAGGUAAUGGCCAAAAACGGAACA .....((((((..(((((((.....))))))).))))))......(((........)))........................... (-12.96 = -13.34 + 0.38)

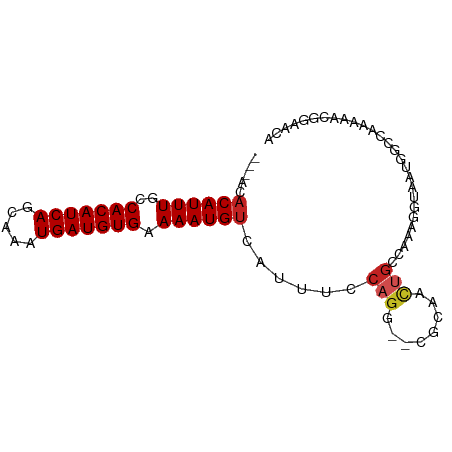
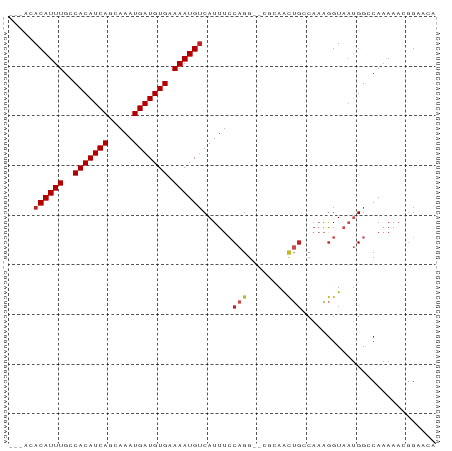
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:06:06 2011