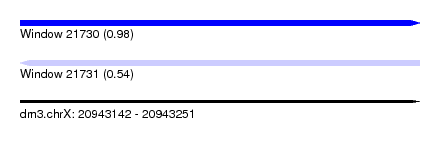

| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,943,142 – 20,943,251 |
| Length | 109 |
| Max. P | 0.977372 |

| Location | 20,943,142 – 20,943,251 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 89.52 |
| Shannon entropy | 0.17301 |
| G+C content | 0.43989 |
| Mean single sequence MFE | -32.26 |
| Consensus MFE | -23.26 |
| Energy contribution | -22.98 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.14 |
| Mean z-score | -3.10 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.97 |
| SVM RNA-class probability | 0.977372 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
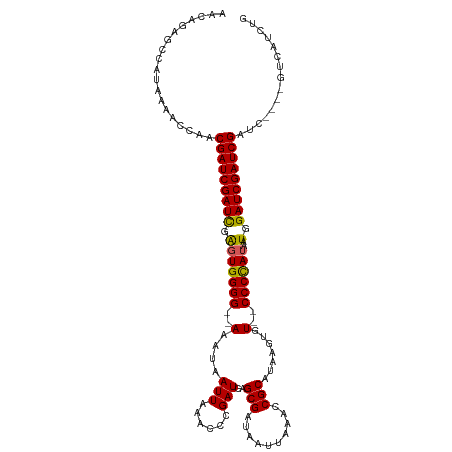
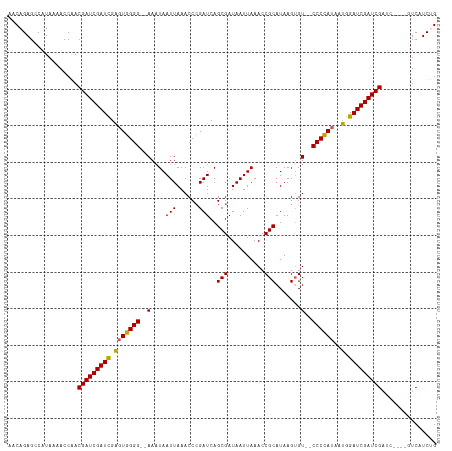
>dm3.chrX 20943142 109 + 22422827 AACAGAGCCAUAAAACCAACGAUCGAUCGAGUGGGG--AAAUAAUUAAACCCGAUCAGCGAUAAUUAAACCGCAUAAGUGU--CCCCAUAAUGGAUCGAUCGAUC----GUCAUCUG ..((((............((((((((((((((((((--(..((((((....((.....)).))))))...(((....))))--))))))......))))))))))----))..)))) ( -31.74, z-score = -3.52, R) >droSim1.chrX 16100638 113 + 17042790 AACAGAGCCAUAAAACCGACGAUCGAUCGAGUGGGG--AAAUAAUUAAACCCGAUCAGCGAUAAUUAAACCGCAUAAGUGU--CCCCAUAAUGGAUCGAUCGAUCGAUCGUCAUCCG .................(((((((((((((..(((.--...........)))((((.(((..........)))....(((.--...)))....))))..)))))))))))))..... ( -35.10, z-score = -3.41, R) >droSec1.super_8 3230861 113 + 3762037 AACAGAGCCAUAAAACCGACGAUCGAUCGAUUGGGG--AAAUAAUUAAACCCGAUCAGCGAUAAUUAAACCGCAUAAGUGU--CCCCAUAAUGGAUCGAUCGAUCGAUCGUCAUCCG .................((((((((((((((((((.--...........)))((((.(((..........)))....(((.--...)))....)))))))))))))))))))..... ( -37.40, z-score = -4.17, R) >droYak2.chrX 20150886 107 + 21770863 AACAGAGCCAUAAAAUCAACGAUCGAUCGAGUGGGGC-AACUAAUUAAACCCGAUCAGCGAUAAUUAAACCGCAUAAGUGU-CCCCCAUAAUGGAUCGAUCG--------UCAUCUG ..((((............((((((((((.(((((((.-.(((.(((......)))..(((..........)))...)))..-.))))))..).)))))))))--------)..)))) ( -28.54, z-score = -2.82, R) >droEre2.scaffold_4690 17514604 108 + 18748788 AACAGAGCCAUACAAUCAACGAUCGAUUGGGUGGGGGGAAAUAAUUAAACC-GAUCAGCGAUAAUUAAACCGCAUAAGAGUGCCCCUAUAAUGGAUCGAUCG--------UCAUCUG ..((((............((((((((((..((((((((...((((((...(-(.....)).))))))..))((((....))))))))))....)))))))))--------)..)))) ( -28.54, z-score = -1.58, R) >consensus AACAGAGCCAUAAAACCAACGAUCGAUCGAGUGGGG__AAAUAAUUAAACCCGAUCAGCGAUAAUUAAACCGCAUAAGUGU__CCCCAUAAUGGAUCGAUCGAUC____GUCAUCUG ...................(((((((((.(((((((.......(((......)))..(((..........)))..........))))))..).)))))))))............... (-23.26 = -22.98 + -0.28)

| Location | 20,943,142 – 20,943,251 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 89.52 |
| Shannon entropy | 0.17301 |
| G+C content | 0.43989 |
| Mean single sequence MFE | -34.72 |
| Consensus MFE | -25.12 |
| Energy contribution | -25.04 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.544944 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
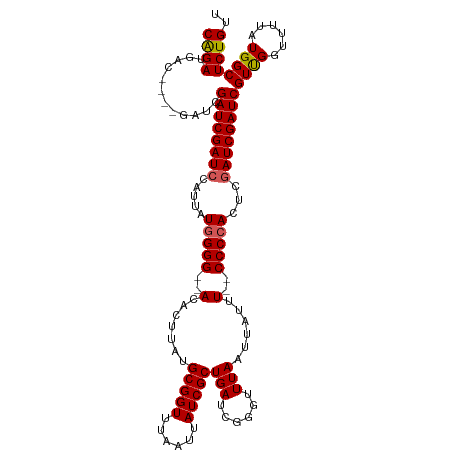
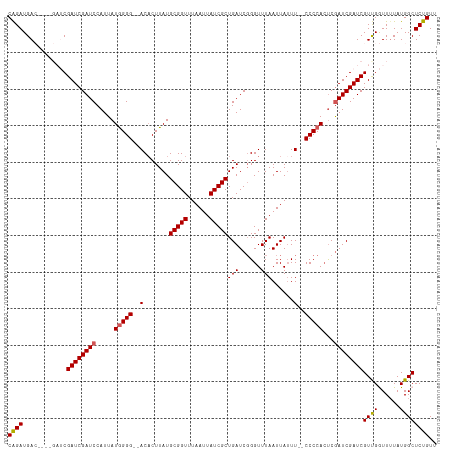
>dm3.chrX 20943142 109 - 22422827 CAGAUGAC----GAUCGAUCGAUCCAUUAUGGGG--ACACUUAUGCGGUUUAAUUAUCGCUGAUCGGGUUUAAUUAUUU--CCCCACUCGAUCGAUCGUUGGUUUUAUGGCUCUGUU ((((.(((----((((((((((((.....((...--.)).....(((((......))))).))))(((...........--.)))....))))))))))).(((....))))))).. ( -34.60, z-score = -2.37, R) >droSim1.chrX 16100638 113 - 17042790 CGGAUGACGAUCGAUCGAUCGAUCCAUUAUGGGG--ACACUUAUGCGGUUUAAUUAUCGCUGAUCGGGUUUAAUUAUUU--CCCCACUCGAUCGAUCGUCGGUUUUAUGGCUCUGUU ((((((((((((((((((..((((.....((...--.)).....(((((......))))).))))(((...........--.)))..))))))))))))))(((....))))))).. ( -39.60, z-score = -2.60, R) >droSec1.super_8 3230861 113 - 3762037 CGGAUGACGAUCGAUCGAUCGAUCCAUUAUGGGG--ACACUUAUGCGGUUUAAUUAUCGCUGAUCGGGUUUAAUUAUUU--CCCCAAUCGAUCGAUCGUCGGUUUUAUGGCUCUGUU (((((((((((((((((((.((((.....((...--.)).....(((((......))))).))))(((...........--.))).)))))))))))))))(((....))))))).. ( -40.40, z-score = -2.74, R) >droYak2.chrX 20150886 107 - 21770863 CAGAUGA--------CGAUCGAUCCAUUAUGGGGG-ACACUUAUGCGGUUUAAUUAUCGCUGAUCGGGUUUAAUUAGUU-GCCCCACUCGAUCGAUCGUUGAUUUUAUGGCUCUGUU (((((((--------(((((((((.....((((((-((......(((((......)))))(((......)))....)))-.)))))...))))))))))))..........)))).. ( -31.30, z-score = -1.44, R) >droEre2.scaffold_4690 17514604 108 - 18748788 CAGAUGA--------CGAUCGAUCCAUUAUAGGGGCACUCUUAUGCGGUUUAAUUAUCGCUGAUC-GGUUUAAUUAUUUCCCCCCACCCAAUCGAUCGUUGAUUGUAUGGCUCUGUU ((((...--------.((((((((....((((((....))))))(((((......))))).))))-)))).............(((..(((((((...)))))))..))).)))).. ( -27.70, z-score = -0.92, R) >consensus CAGAUGAC____GAUCGAUCGAUCCAUUAUGGGG__ACACUUAUGCGGUUUAAUUAUCGCUGAUCGGGUUUAAUUAUUU__CCCCACUCGAUCGAUCGUUGGUUUUAUGGCUCUGUU ((((............((((((((.....(((((..........(((((......)))))(((......))).........)))))...))))))))((((......)))))))).. (-25.12 = -25.04 + -0.08)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:05:33 2011