
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,927,355 – 20,927,454 |
| Length | 99 |
| Max. P | 0.649282 |

| Location | 20,927,355 – 20,927,454 |
|---|---|
| Length | 99 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 66.20 |
| Shannon entropy | 0.61505 |
| G+C content | 0.34452 |
| Mean single sequence MFE | -16.86 |
| Consensus MFE | -7.07 |
| Energy contribution | -6.87 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.58 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.649282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20927355 99 - 22422827 ------AAAUUAUUUUGAGGCGAAAGCAG--AAAUCGAAGAAAAAAUAAACAAAAACACUUAAU-CACUC----AAUGGCCAUUUA-AAUGGGUUAGGAAAUGGUCGACAUCA ------........(((((((....)).(--....)............................-..)))----))(((((((((.-...........)))))))))...... ( -16.30, z-score = -1.27, R) >droAna3.scaffold_13334 274674 104 + 1562580 AAAAAACGAUGUUUUUGGAGCGAAAGCAGCAAAAAAAAAAAGAAAAGACAAGAAAUCACUUAAU-CACUCGGUUGUUGGUCAUGUA-AAUGGGUUAGGAAAUGGCC------- ....((((((.((((((..((....))..))))))..............(((......)))...-......))))))((((((...-.............))))))------- ( -15.99, z-score = 0.13, R) >droEre2.scaffold_4690 17499305 86 - 18748788 ------AAAUUAUUUUGGGGCGAAAGCAG---AAAUGAUAAACAAAAAUAAAAA--CACUUAAU-CACUC----AUUGGCCAUGUA-AAUGGGUUAGGAAAUG---------- ------..((((((((...((....)).)---)))))))...............--..((((..-.((((----(((.((...)).-))))))))))).....---------- ( -16.30, z-score = -2.27, R) >droYak2.chrX 20135633 92 - 21770863 ------AAAUUAUUUCGAGGCGAAAGCAG---AAAUAAAAAAAAAA----------CACUUAAU-CACUCAUUGGUCAUGUAUGUA-AAUGGGUUAGGAAAUGGUCGACAUAA ------...(((((((...((....)).)---))))))........----------..((((..-.(((((((...((....))..-)))))))))))..(((.....))).. ( -18.00, z-score = -2.49, R) >droSec1.super_8 3216147 99 - 3762037 ------AGAUUAUUUUGAGGCGAAAGCAG--AAAUCGAAGAAAAAAUAAACAAAAACACUUAAU-CACUC----AAUGGCCAUUUA-AAUGGGUUAGGAAAUGGUCGACCUCA ------.((((((((((((((....)).(--....)............................-..)))----..(((((.....-....))))).)))))))))....... ( -16.80, z-score = -0.69, R) >droSim1.chrX_random 621757 99 - 5698898 ------AGAUUAUUUUGAGGCGAAAGCAG--AAAUCGAAAAAAAAAUAAACAAAAACACUUAAU-CACUC----AAUGGCCAUUUA-AAUGGGUUAGGAAAUGGUCGACAUCA ------.(((....(((((((....)).(--....)............................-..)))----))(((((((((.-...........)))))))))..))). ( -17.00, z-score = -1.54, R) >droVir3.scaffold_12970 7673284 87 - 11907090 ------UUUUUUUAUUGGAGCGAAAGCA----AAAUGCAGCAGACCAAUAAAAUUACUCUUAGCGUACCC----AAAGGCCCCGUAGAGCAGGCCAACUUC------------ ------...(((((((((.((....)).----.....(....).))))))))).................----...((((..(.....).))))......------------ ( -17.60, z-score = -0.68, R) >consensus ______AAAUUAUUUUGAGGCGAAAGCAG__AAAACGAAAAAAAAAUAAAAAAAAACACUUAAU_CACUC____AAUGGCCAUGUA_AAUGGGUUAGGAAAUGGUCGAC_U_A ............((((((.((....))....................................................((((.....)))).)))))).............. ( -7.07 = -6.87 + -0.20)


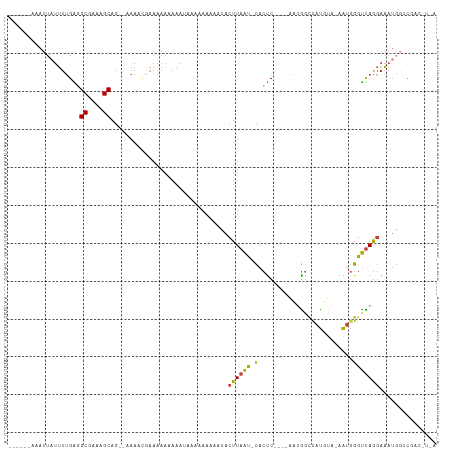
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:05:28 2011