
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,736,216 – 20,736,332 |
| Length | 116 |
| Max. P | 0.981478 |

| Location | 20,736,216 – 20,736,332 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 80.35 |
| Shannon entropy | 0.26699 |
| G+C content | 0.61037 |
| Mean single sequence MFE | -38.32 |
| Consensus MFE | -22.49 |
| Energy contribution | -23.72 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.805463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
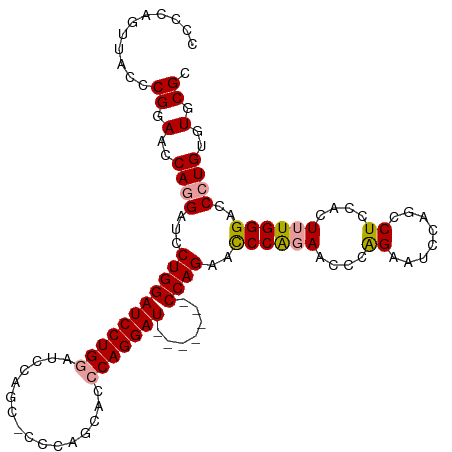
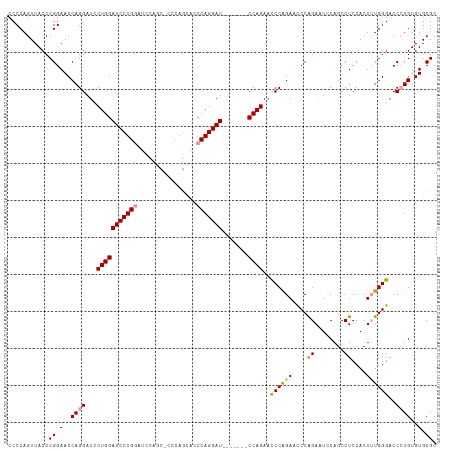
>dm3.chrX 20736216 116 + 22422827 CCCCAGUUACCCGGAACCAGGAUUCUGGAUCCUGGAUCCAGCGCCCAGUACCCAGGAUUCAGAAUCCAGAACCCAGAACCCAGAAUCCAGCCUCCACUUUGGGACCCUGUGUGCGC .....((.((.(((.....((((((((((((((((.....((.....))..))))))))))))))))....((((((....((........))....))))))...))).)))).. ( -38.10, z-score = -2.31, R) >droEre2.scaffold_4690 17319215 95 + 18748788 CCCCAGUUACCCGGAACCAGGAUCCUGGAUCCUG--------------CGCCCAGGAU-------CCAGAAUCCGGAAUCCGGACUACAGCCUCCACUGUGGGACCAUGUGUGCGC .(((......(((((.((.((((.((((((((((--------------....))))))-------)))).))))))..)))))...((((......)))))))............. ( -40.80, z-score = -3.25, R) >droSim1.chrX_random 382597 109 + 5698898 CCCCAGUUACCCGGAACCAGGAUCCUGGAUCCUGGAUCCAGCACCCAGCACCCAGGAU-------CCAGAGCCCAGAACCCAGAAUCCAGCCUCCACUUUGGGACCCUGUGUGCGC ............(((.((((((((...)))))))).)))........((((.((((.(-------((((((...((...............))...)))))))..)))).)))).. ( -36.06, z-score = -1.77, R) >consensus CCCCAGUUACCCGGAACCAGGAUCCUGGAUCCUGGAUCCAGC_CCCAGCACCCAGGAU_______CCAGAACCCAGAACCCAGAAUCCAGCCUCCACUUUGGGACCCUGUGUGCGC ............(((.((((((((...)))))))).)))........((((.((((............(....)....((((((.............))))))..)))).)))).. (-22.49 = -23.72 + 1.23)

| Location | 20,736,216 – 20,736,332 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 80.35 |
| Shannon entropy | 0.26699 |
| G+C content | 0.61037 |
| Mean single sequence MFE | -51.67 |
| Consensus MFE | -38.27 |
| Energy contribution | -39.14 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.981478 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
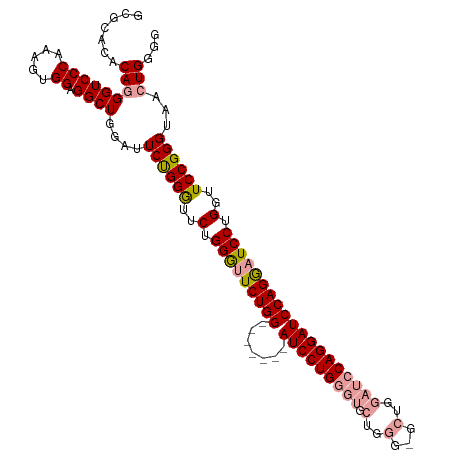
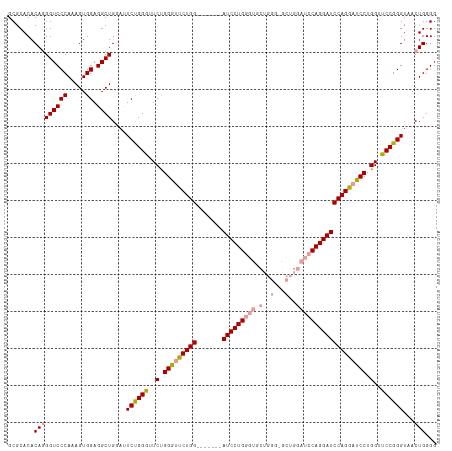
>dm3.chrX 20736216 116 - 22422827 GCGCACACAGGGUCCCAAAGUGGAGGCUGGAUUCUGGGUUCUGGGUUCUGGAUUCUGAAUCCUGGGUACUGGGCGCUGGAUCCAGGAUCCAGAAUCCUGGUUCCGGGUAACUGGGG .......(((...(((..(((....)))(((..(.((((((((((((((((((((.(..(((........)))..).)))))))))))))))))))).)..))))))...)))... ( -53.30, z-score = -2.53, R) >droEre2.scaffold_4690 17319215 95 - 18748788 GCGCACACAUGGUCCCACAGUGGAGGCUGUAGUCCGGAUUCCGGAUUCUGG-------AUCCUGGGCG--------------CAGGAUCCAGGAUCCUGGUUCCGGGUAACUGGGG .............((((((((....)))))..((((((..(((((((((((-------((((((....--------------))))))))))))))).)).)))))).....))). ( -48.60, z-score = -3.30, R) >droSim1.chrX_random 382597 109 - 5698898 GCGCACACAGGGUCCCAAAGUGGAGGCUGGAUUCUGGGUUCUGGGCUCUGG-------AUCCUGGGUGCUGGGUGCUGGAUCCAGGAUCCAGGAUCCUGGUUCCGGGUAACUGGGG .......((((((((......((((.(..((........))..).))))((-------(((((((((.(........).))))))))))).))))))))..(((((....))))). ( -53.10, z-score = -2.18, R) >consensus GCGCACACAGGGUCCCAAAGUGGAGGCUGGAUUCUGGGUUCUGGGUUCUGG_______AUCCUGGGUGCUGGG_GCUGGAUCCAGGAUCCAGGAUCCUGGUUCCGGGUAACUGGGG .......(((((((((.....)).))))....((((((..(.(((((((((.......(((((((((.(........).)))))))))))))))))).)..))))))...)))... (-38.27 = -39.14 + 0.87)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:04:56 2011