
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 12,185,058 – 12,185,148 |
| Length | 90 |
| Max. P | 0.961460 |

| Location | 12,185,058 – 12,185,148 |
|---|---|
| Length | 90 |
| Sequences | 11 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 75.86 |
| Shannon entropy | 0.45415 |
| G+C content | 0.33855 |
| Mean single sequence MFE | -17.05 |
| Consensus MFE | -8.67 |
| Energy contribution | -8.67 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.961460 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 12185058 90 + 23011544 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA-----------GCUACGA----ACAAGUGAGG---AAAAAUCAUAAAAAUUU-GCGCUACAAAUAAACAA-----ACCAAUAAAAG --((((((((....))))))))...((((.-----------((.....----....((((..---.....))))........-))))))...........-----........... ( -15.13, z-score = -2.39, R) >droSim1.chr2L 11985783 90 + 22036055 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA-----------GCUACGA----ACAAGUGAGG---AAAAAUCAUAAAAAUUU-GCGCUACAAAUAAACAA-----ACCAAUAAAAG --((((((((....))))))))...((((.-----------((.....----....((((..---.....))))........-))))))...........-----........... ( -15.13, z-score = -2.39, R) >droSec1.super_16 380429 90 + 1878335 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA-----------GCUACGA----ACAAGUGAGG---AAAAAUCAUAAAAAUUU-GCGCUACAAAUAAACAA-----UCCAAUAAAAG --((((((((....))))))))...((((.-----------((.....----....((((..---.....))))........-))))))...........-----........... ( -15.13, z-score = -2.21, R) >droYak2.chr2L 8615467 93 + 22324452 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA-----------GCUACGAU-GAACAAGUGAGG---AAAAAUCAUAAAAAUUU-GUGCUACAAAUAAACAA-----ACCAAUAAAAG --((((((((....))))))))(((((..(-----------((.((((.-......((((..---.....))))......))-))))).....)))))..-----........... ( -17.62, z-score = -2.74, R) >droEre2.scaffold_4929 13414041 90 - 26641161 --UGAUUUCCCCAUGGAAAUCAGUUUAGC-----------CACUACG----AGCAAGUGAGG---AAAAAUCAUAAAAAUGU-GUGCUACAAAUAAACAA-----ACCAAUAAAAG --((((((((....))))))))...((((-----------(((....----.....((((..---.....))))......))-).))))...........-----........... ( -16.06, z-score = -2.32, R) >droAna3.scaffold_12916 2982411 89 - 16180835 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA-----------G-UACGU----ACCAGAAAGG---AAAAAUCGUAAAAAUUU-GUGCUACAAAUAAACAA-----ACCAAUAAAAG --((((((((....))))))))(((((..(-----------(-(((..----.((.....))---..((((.......))))-))))).....)))))..-----........... ( -15.20, z-score = -2.54, R) >droPer1.super_1 2810361 93 - 10282868 --UGAUUUCCCCAUGGAAAUCAGUUUAGAA-------------CGCA-----GCUAAAGAGC-AAAAAAAUCAUAAAAAUUU-GUGCUACAAAUAAACAAAUAAAACCAAUAAAA- --((((((((....))))))))(((((...-------------.(((-----(((....)))-........((........)-))))......))))).................- ( -14.80, z-score = -2.95, R) >droWil1.scaffold_180708 9865452 113 + 12563649 GUUGAUUUCCAUGGGGAAAUCAGUUUAGUAGCAGCUACAGCACCACAUCCAAGCAGUUGGGGCAUAAAAAUCAUAAAAAUUU-GUGCUACAAAUAAACAAG--AAACCAAUAAAAG ..((((((((....))))))))(((((((((((((..((((..............))))..))........((........)-)))))))...)))))...--............. ( -24.94, z-score = -2.23, R) >droVir3.scaffold_12963 17245697 84 + 20206255 --UGAUUUCCCCAUGGAAAUCAGUUUAGCA----------------------G---UUGGGC-UA-AAAAUCAUAAAA-UCUAGCGCUACCAAUAAAAACA--CGACCGAUGAAAA --((((((((....))))))))...((((.----------------------(---((((..-(.-..........).-.)))))))))............--............. ( -15.10, z-score = -1.35, R) >droMoj3.scaffold_6500 22287217 84 + 32352404 --UGAUUUCCCAAUGGAAAUCAGUUUAGCA----------------------G---UUGGGC-UA-AAAAUCAUAAAA-UCUAGCGCUACAAAUAAAAAUG--CGACCGAUAAAAA --((((((((....)))))))).((((((.----------------------.---....))-))-)).........(-((...(((.............)--))...)))..... ( -17.82, z-score = -2.47, R) >droGri2.scaffold_15252 14675543 88 + 17193109 --UGAUUUCCCCAUGGAAAUCAGUUUAGCU----------------------GCGCUUGGGC-UGGAAAAUCAUAAAAGUCUUGUGAUACAAAUAAAAAC---CAUGCGAUGAAAC --((((((((....))))))))((((....----------------------.(((.(((..-......(((((((.....)))))))...........)---)).)))...)))) ( -20.61, z-score = -1.87, R) >consensus __UGAUUUCCCCAUGGAAAUCAGUUUAGCA___________GCUACG_____GCAAGUGAGG___AAAAAUCAUAAAAAUUU_GUGCUACAAAUAAACAA_____ACCAAUAAAAG ..((((((((....)))))))).............................................................................................. ( -8.67 = -8.67 + -0.00)
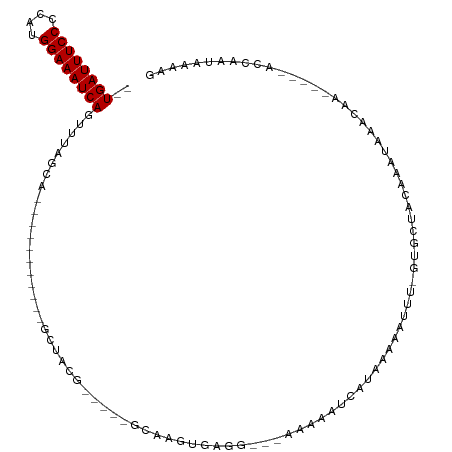


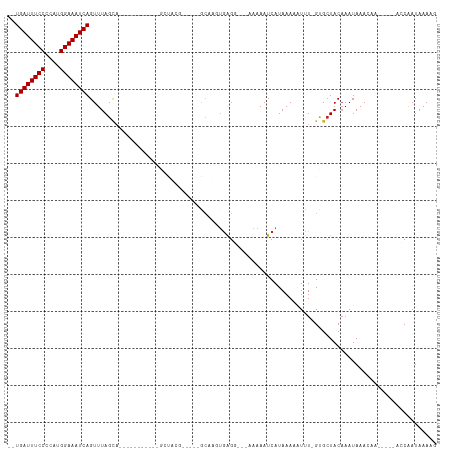
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:34:32 2011