
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,478,475 – 20,478,576 |
| Length | 101 |
| Max. P | 0.973735 |

| Location | 20,478,475 – 20,478,576 |
|---|---|
| Length | 101 |
| Sequences | 10 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 84.33 |
| Shannon entropy | 0.34044 |
| G+C content | 0.44809 |
| Mean single sequence MFE | -29.20 |
| Consensus MFE | -21.61 |
| Energy contribution | -21.71 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.973735 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20478475 101 + 22422827 CAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGA-CCUGUCCGUUCGUCCACC ..((.(((((..(.(((((((.....((((((((....)))))))).....))))))).)((((.(((((((....)))))).-).)))).).)))).)).. ( -35.00, z-score = -3.78, R) >droSim1.chrX_random 5342895 101 + 5698898 CAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGA-CCUGUCCGUCCGUCCGCC ...(.(((((..(.(((((((.....((((((((....)))))))).....))))))).)((((.(((((((....)))))).-).)))).)).))).)... ( -34.30, z-score = -3.18, R) >droSec1.super_8 2720163 101 + 3762037 CAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGA-CCUGUCCGUCCGUCCGCC ...(.(((((..(.(((((((.....((((((((....)))))))).....))))))).)((((.(((((((....)))))).-).)))).)).))).)... ( -34.30, z-score = -3.18, R) >droYak2.chrX 19708092 97 + 21770863 CAUGUACGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGAA-CUUGUCCGUCCACC---- .......(((..(.(((((((.....((((((((....)))))))).....))))))).)((((...(((((....)))))..-..)))).)))....---- ( -29.50, z-score = -2.92, R) >droEre2.scaffold_4690 17079736 97 + 18748788 CAUGUAGGACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGA-CUUGUCCAUCCACC---- ......(((...(.(((((((.....((((((((....)))))))).....))))))).)((((..((((((....)))))).-..))))..)))...---- ( -34.50, z-score = -3.83, R) >droAna3.scaffold_12903 557329 94 - 802071 CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACUCAUAGA-GUCCUUUUUCG------- .........((((((((((((.....((((((((....)))))))).....)))))))...)))))....((((.(.....).-)))).......------- ( -23.70, z-score = -1.87, R) >dp4.chrXL_group1e 9112703 97 + 12523060 CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUAGAUACACACACA-CACACACACGCACU---- .........((((((((((((.....((((((((....)))))))).....)))))))...))))).................-..............---- ( -20.40, z-score = -0.86, R) >droVir3.scaffold_12970 4771610 97 + 11907090 CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUU-CGACAUGCCAUGGAUACUCUGAGAGUGUAGGUGUGCGCU---- ...(((((.((((((((((((.....((((((((....)))))))).....)))))))-..)))))((....((((((...)))))).)))))))...---- ( -30.80, z-score = -1.60, R) >droMoj3.scaffold_6473 4215153 99 + 16943266 CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCAACAUGCCAUGGAUAUUGUGAGAGUUUGAGC-UGCCCUCU-- ((((.....((((((((((((.....((((((((....)))))))).....)))))))...))))).))))........(((.((......-.)).))).-- ( -24.90, z-score = -0.53, R) >droGri2.scaffold_15081 3725140 92 - 4274704 CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCAACAUGCCAUGGAUACGCUGCGACUAUGUGU-G--------- ((((.....((((((((((((.....((((((((....)))))))).....)))))))...))))).))))..(((((.........))))-)--------- ( -24.60, z-score = -0.44, R) >consensus CAUGUACAACAUGUAAAUUGUUUCAAUUUGCGGCAUUUGCCGUAAAGCUGCGCAAUUUUCGACAUGCCAUGGAUACCCAUGGA_CUUGUCCGUCCGCC____ ((((.....((((((((((((.....((((((((....)))))))).....)))))))...))))).))))............................... (-21.61 = -21.71 + 0.10)
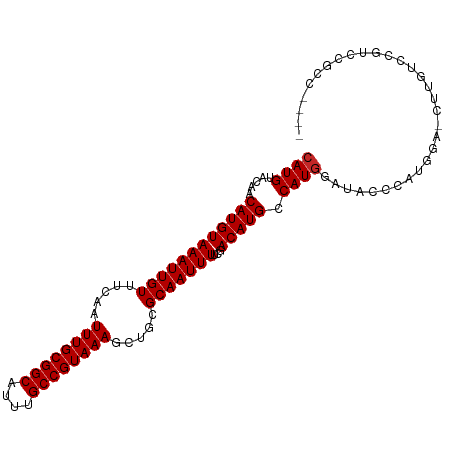


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:04:15 2011