
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,442,771 – 20,442,897 |
| Length | 126 |
| Max. P | 0.989495 |

| Location | 20,442,771 – 20,442,897 |
|---|---|
| Length | 126 |
| Sequences | 11 |
| Columns | 144 |
| Reading direction | forward |
| Mean pairwise identity | 78.07 |
| Shannon entropy | 0.43728 |
| G+C content | 0.48663 |
| Mean single sequence MFE | -41.41 |
| Consensus MFE | -23.89 |
| Energy contribution | -23.72 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.10 |
| SVM RNA-class probability | 0.982399 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20442771 126 + 22422827 ----------GAAAACGGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGC-UAGACGAAAAUUCCCUCGCUCCAAAAACU------ ----------......(((((((...((..((((....)))).))(((((((...)))))))....(((((((...((((((((......))))))))......-(..-..)..)))))))...))))))).......------ ( -42.00, z-score = -2.42, R) >droPer1.super_22 549006 142 - 1688296 GAGAGCGGGAGAGAGGACAUGGGAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGU-UAGACGAAAAUUCCCUCGUGCUCGUGCCACAUUCU .((((.(((...(((.((..(((((.((..((((....)))).))(((((((...)))))))(((((..((((...((((((((......))))))))))))))-)))-..........)))))..)).)))...)).).)))) ( -48.00, z-score = -1.66, R) >dp4.chrXL_group1e 9069155 142 + 12523060 GAGAGUGGGGGAGAGGACAUGGGAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGU-UAGACGAAAAUUCCCUCGUGCUCGUGCCACAUUCU .((((((((.(.(((.((..(((((.((..((((....)))).))(((((((...)))))))(((((..((((...((((((((......))))))))))))))-)))-..........)))))..)).))).).)).)))))) ( -52.30, z-score = -2.83, R) >droAna3.scaffold_12903 520364 123 - 802071 ------------GAAAUGGGAGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUUUCGC-UAGACGAAAAUUUUAUCGCUC-CAAGCU------- ------------(((((..(((((..(((.((((....))))...(((((((...))))))).)))..)))))...((((((((......))))))))...)))))((-(.(((((........))).))-..))).------- ( -37.50, z-score = -1.54, R) >droEre2.scaffold_4690 17043273 126 + 18748788 ----------GAAAACGGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGC-UAGACGAAAAUUCCCUCGCUCCAAAAACU------ ----------......(((((((...((..((((....)))).))(((((((...)))))))....(((((((...((((((((......))))))))......-(..-..)..)))))))...))))))).......------ ( -42.00, z-score = -2.42, R) >droYak2.chrX 19670324 126 + 21770863 ----------GAAAACGGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCCCGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGC-UAGACGAAAAUUCCCUCGCUCCAAAAACU------ ----------......(((((((..((...((((....))))((((((((((...)))))))..(((..((((...((((((((......))))))))))))))-)))-)............))))))))).......------ ( -41.70, z-score = -2.03, R) >droSec1.super_8 2673594 126 + 3762037 ----------GAAAACGGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGC-UAGACGAAAAUUCCCUCGCUCCAAAAACU------ ----------......(((((((...((..((((....)))).))(((((((...)))))))....(((((((...((((((((......))))))))......-(..-..)..)))))))...))))))).......------ ( -42.00, z-score = -2.42, R) >droSim1.chrX 15671353 126 + 17042790 ----------GAAAACGGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA-CGC-UAGACGAAAAUUCCCUCGCUCCAAAAACU------ ----------......(((((((...((..((((....)))).))(((((((...)))))))....(((((((...((((((((......))))))))......-(..-..)..)))))))...))))))).......------ ( -42.00, z-score = -2.42, R) >droVir3.scaffold_12970 4729855 119 + 11907090 -------------AAAAACCGAGCAGCCUCAUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACG--AAAAUACGUAAACGAAAACUGGAUGCCAAAACU---------- -------------.......(.((((((((((.....)))).)))(((((((...)))))))(((((.(((((...((((((((......)))))))))--)))))))))..............))))......---------- ( -35.30, z-score = -2.33, R) >droMoj3.scaffold_6473 4163737 118 + 16943266 ----------------AACCGAGCAGCCUCAUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAGAAAAUACGUAAACGAAAACUCGAUGCCAAAAAC---------- ----------------...((((..(((((((.....)))).)))(((((((...)))))))(((((.(((((.((.(((((((......))))))))).)))))))))).........))))...........---------- ( -38.30, z-score = -3.30, R) >droGri2.scaffold_15081 3688391 112 - 4274704 -------------AAUAACCGAGCAGCCCCAUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGAUUUUUCGCAAAUACGGAAAACGAGAAAAUACGUAAACGAAAACUCGCU------------------- -------------......((((..(((...(((....))).)))(((((((...)))))))(((((.(((((.((.(((((((......))))))))).)))))))))).........))))..------------------- ( -34.40, z-score = -2.54, R) >consensus __________GAAAACAGGGCGAAAGGCAGGUCAUGAGUGACGGCGCCGCUGCGUCAGCGGCAUGUAAUUUUCACGGUUUUUCGCAAAUACGAAAAACGAAAUA_CGC_UAGACGAAAAUUCCCUCGCUCCAAAAACU______ ...................................((((..((..(((((((...)))))))..............((((((((......))))))))...............))...))))...................... (-23.89 = -23.72 + -0.17)


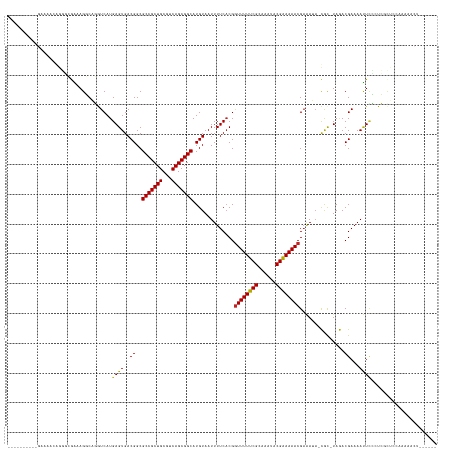
| Location | 20,442,771 – 20,442,897 |
|---|---|
| Length | 126 |
| Sequences | 11 |
| Columns | 144 |
| Reading direction | reverse |
| Mean pairwise identity | 78.07 |
| Shannon entropy | 0.43728 |
| G+C content | 0.48663 |
| Mean single sequence MFE | -40.51 |
| Consensus MFE | -26.45 |
| Energy contribution | -25.86 |
| Covariance contribution | -0.59 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.989495 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20442771 126 - 22422827 ------AGUUUUUGGAGCGAGGGAAUUUUCGUCUA-GCG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCGCCCCGUUUUC---------- ------.......((.(((((((((((((((....-.).-......(((((((((....)))))))))...)))))))....(((((((...)))))))((.((((....))))..))))))))).))......---------- ( -41.20, z-score = -2.26, R) >droPer1.super_22 549006 142 + 1688296 AGAAUGUGGCACGAGCACGAGGGAAUUUUCGUCUA-ACG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUCCCAUGUCCUCUCUCCCGCUCUC .((..((((...(((.(((.(((((.....(((..-...-......(((((((((....))))))))).((((........((((((((...))))))))...))))....))).....))))).))).)))....))))..)) ( -44.20, z-score = -2.46, R) >dp4.chrXL_group1e 9069155 142 - 12523060 AGAAUGUGGCACGAGCACGAGGGAAUUUUCGUCUA-ACG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUCCCAUGUCCUCUCCCCCACUCUC .((..((((...(((.(((.(((((.....(((..-...-......(((((((((....))))))))).((((........((((((((...))))))))...))))....))).....))))).))).)))....))))..)) ( -44.60, z-score = -3.04, R) >droAna3.scaffold_12903 520364 123 + 802071 -------AGCUUG-GAGCGAUAAAAUUUUCGUCUA-GCGAAAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCUCCCAUUUC------------ -------.(((.(-((.(((........)))))))-))(((((...(((((((((....))))))))).(.((((.......(((((((...)))))))((.((((....))))..)).)))).)..)))))------------ ( -35.40, z-score = -2.29, R) >droEre2.scaffold_4690 17043273 126 - 18748788 ------AGUUUUUGGAGCGAGGGAAUUUUCGUCUA-GCG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCGCCCCGUUUUC---------- ------.......((.(((((((((((((((....-.).-......(((((((((....)))))))))...)))))))....(((((((...)))))))((.((((....))))..))))))))).))......---------- ( -41.20, z-score = -2.26, R) >droYak2.chrX 19670324 126 - 21770863 ------AGUUUUUGGAGCGAGGGAAUUUUCGUCUA-GCG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGGGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCGCCCCGUUUUC---------- ------.......((.(((((((((((((((....-.).-......(((((((((....)))))))))...)))))))....(((((((...)))))))((.((((....))))..))))))))).))......---------- ( -41.20, z-score = -1.69, R) >droSec1.super_8 2673594 126 - 3762037 ------AGUUUUUGGAGCGAGGGAAUUUUCGUCUA-GCG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCGCCCCGUUUUC---------- ------.......((.(((((((((((((((....-.).-......(((((((((....)))))))))...)))))))....(((((((...)))))))((.((((....))))..))))))))).))......---------- ( -41.20, z-score = -2.26, R) >droSim1.chrX 15671353 126 - 17042790 ------AGUUUUUGGAGCGAGGGAAUUUUCGUCUA-GCG-UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCGCCCCGUUUUC---------- ------.......((.(((((((((((((((....-.).-......(((((((((....)))))))))...)))))))....(((((((...)))))))((.((((....))))..))))))))).))......---------- ( -41.20, z-score = -2.26, R) >droVir3.scaffold_12970 4729855 119 - 11907090 ----------AGUUUUGGCAUCCAGUUUUCGUUUACGUAUUUU--CGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGAUGAGGCUGCUCGGUUUUU------------- ----------......(((((..((((((((....).......--.(((((((((....)))))))))...)))))))..)))))(((((.((((((((...))).(((.....)))))))))))))....------------- ( -37.90, z-score = -2.15, R) >droMoj3.scaffold_6473 4163737 118 - 16943266 ----------GUUUUUGGCAUCGAGUUUUCGUUUACGUAUUUUCUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGAUGAGGCUGCUCGGUU---------------- ----------.........(((((((..........((((((((.((((((((((....)))))))).)).))))).)))..(((((((...)))))))(((.((((....).)))))).))))))).---------------- ( -40.30, z-score = -2.66, R) >droGri2.scaffold_15081 3688391 112 + 4274704 -------------------AGCGAGUUUUCGUUUACGUAUUUUCUCGUUUUCCGUAUUUGCGAAAAAUCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGAUGGGGCUGCUCGGUUAUU------------- -------------------..(((((..((......((((((((.((((((.(((....))).)))).)).))))).)))..(((((((...))))))).((((((....))))))))..)))))......------------- ( -37.20, z-score = -1.98, R) >consensus ______AGUUUUUGGAGCGAGGGAAUUUUCGUCUA_GCG_UAUUUCGUUUUUCGUAUUUGCGAAAAACCGUGAAAAUUACAUGCCGCUGACGCAGCGGCGCCGUCACUCAUGACCUGCCUUUCCCCCCGUUUUC__________ .......................((((((((......)........(((((((((....)))))))))...)))))))....(((((((...)))))))...((((....)))).............................. (-26.45 = -25.86 + -0.59)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:04:12 2011