
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,357,794 – 20,357,920 |
| Length | 126 |
| Max. P | 0.716575 |

| Location | 20,357,794 – 20,357,889 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 62.41 |
| Shannon entropy | 0.58360 |
| G+C content | 0.40555 |
| Mean single sequence MFE | -20.25 |
| Consensus MFE | -8.27 |
| Energy contribution | -8.65 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.628423 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
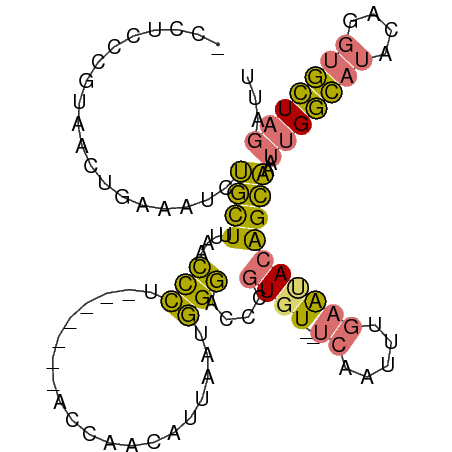
>dm3.chrX 20357794 95 + 22422827 -CCUUUGUGAACUGAAAUCUGCUUUAACCCU-------ACCAACAUUAAUGGGACCCGUGUGUUCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCUAGAUU -..............(((((((.....(((.-------............)))(((.((((((.(((((((.......))))))))))))).))))).))))) ( -24.02, z-score = -1.92, R) >droSim1.chrX 15613801 100 + 17042790 -CCUCCCGUUACUGAAAUCUGCUUUAACCCUACAACCUACCAACAUUAAUGGGACCCGUGU--UCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCUAGAUU -..((((((((.((.............................)).))))))))...((((--((.....))))))......(((((((....)))))))... ( -20.25, z-score = -1.17, R) >droSec1.super_8 2552169 93 + 3762037 -CCUCCCGUUACUGAAAUCUGCUUUAACCCU-------ACCAACAUUAAUGGGACCCGUGU--UCAAUUUGAACACAGCAAAUUGGCAUACAGGUGCUAGAUU -..((((((((.((.................-------.....)).))))))))...((((--((.....))))))......(((((((....)))))))... ( -22.45, z-score = -2.03, R) >droWil1.scaffold_180702 1577035 86 - 4511350 UAAAAAGUCAAGGAAAAUGCAUUUUGAUUGC--------ACACUGUUGACUGAGGCUAUGU---GAAUGUAAGGA-GGUGGACAGAUAGAUAGAGAUU----- ......(((........((((.......)))--------)..(((((.(((...((.....---....)).....-))).)))))...))).......----- ( -14.30, z-score = -0.59, R) >consensus _CCUCCCGUAACUGAAAUCUGCUUUAACCCU_______ACCAACAUUAAUGGGACCCGUGU__UCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCUAGAUU ...............(((((((.....(((....................)))(((.((((...(((((((.......)))))))..)))).))))).))))) ( -8.27 = -8.65 + 0.38)


| Location | 20,357,827 – 20,357,920 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 67.55 |
| Shannon entropy | 0.51087 |
| G+C content | 0.38018 |
| Mean single sequence MFE | -23.40 |
| Consensus MFE | -7.85 |
| Energy contribution | -10.47 |
| Covariance contribution | 2.63 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.716575 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
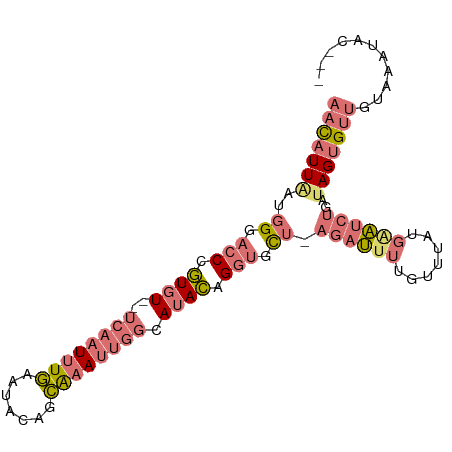
>dm3.chrX 20357827 93 + 22422827 AACAUUAAUGGGACCCGUGUGUUCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCU-AGAUUUUGUUUAUGAAUCUCAAAGUGUUGUAAAUAC--- ((((((..(((.(((.((((((.(((((((.......))))))))))))).))).))-)(((((.......)))))....))))))........--- ( -25.40, z-score = -2.65, R) >droSim1.chrX 15613841 91 + 17042790 AACAUUAAUGGGACCCGUGU--UCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCU-AGAUUUUGUUUAUGGAUCUGAUAGUGUUGUAAAUAC--- (((((((.(.(((.((((((--((.....))))))(((((((((((((....)))))-))..))))))...)).))).))))))))........--- ( -25.00, z-score = -2.58, R) >droSec1.super_8 2552202 91 + 3762037 AACAUUAAUGGGACCCGUGU--UCAAUUUGAACACAGCAAAUUGGCAUACAGGUGCU-AGAUUUUGUUUAUGGAUCUGAUAGUGUUGUAAAUAC--- (((((((.(.(((.((((((--((.....))))))(((((((((((((....)))))-))..))))))...)).))).))))))))........--- ( -26.90, z-score = -3.02, R) >droWil1.scaffold_180702 1577068 93 - 4511350 ACUGUUGACUGAGGCUAUGU---GAAUGUAAGGA-GGUGGACAGAUAGAUAGAGAUUUAGAGAGAGAGCAAGACAGAGCUAGAGUGGGCAGCACAAC .(((((.(((..((((.(((---...(((.....-(.....)...(((((....)))))........)))..))).))))..))).)))))...... ( -16.30, z-score = -0.75, R) >consensus AACAUUAAUGGGACCCGUGU__UCAAUUUGAAUACAGCAAAUUGGCAUACAGGUGCU_AGAUUUUGUUUAUGAAUCUGAUAGUGUUGUAAAUAC___ (((((((.(((((.(((((....(((((((.......)))))))((((....))))............))))).))))))))))))........... ( -7.85 = -10.47 + 2.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:03:57 2011