
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,241,416 – 20,241,525 |
| Length | 109 |
| Max. P | 0.789157 |

| Location | 20,241,416 – 20,241,525 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 70.56 |
| Shannon entropy | 0.52326 |
| G+C content | 0.50272 |
| Mean single sequence MFE | -24.70 |
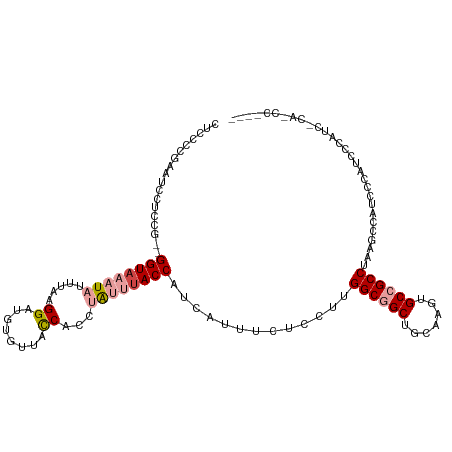
| Consensus MFE | -15.23 |
| Energy contribution | -16.28 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.02 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.789157 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
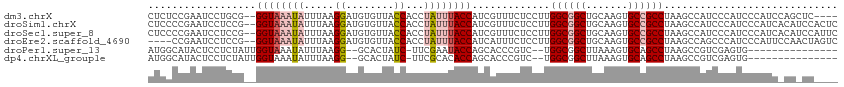
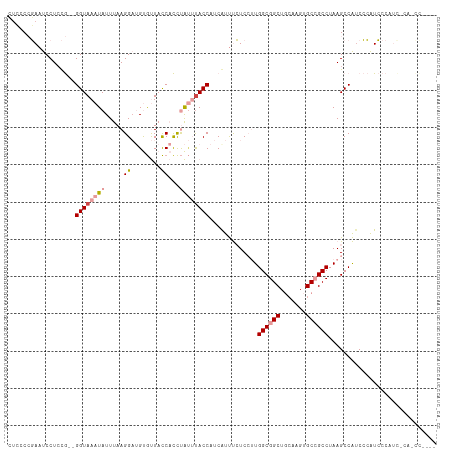
>dm3.chrX 20241416 109 + 22422827 CUCUCCGAAUCCUGCG--GGUAAAUAUUUAAGGAUGUGUUACCACCUAUUUACCAUCGUUUCUCCUUGGCGGCUGCAAGUGCCGCCUAAGCCAUCCCAUCCCAUCCAGCUC---- .............((.--(((((((.....(((.((......))))))))))))..........(((((((((.......)))))).))).................))..---- ( -23.70, z-score = -0.52, R) >droSim1.chrX 15516596 113 + 17042790 CUCCCCGAAUCCUCCG--GGUAAAUAUUUAAGGAUGUGUUACCACCUAUUUACCAUCGUUUCUCCUUGGCGGCUGCAAGUGCCGCCUAAGCCAUCCCAUCCCAUCACAUCCACUC ...((((.......))--))...........(((((((.............((....)).....(((((((((.......)))))).)))..............))))))).... ( -27.90, z-score = -2.76, R) >droSec1.super_8 2428469 113 + 3762037 CUCCCCGAAUCCUCCG--GGUAAAUAUUUAAGGAUGUGUUACCACCUAUUUACCAUCGUUUCUCCUUGGCGGCUGCAAGUGCCGCCUAAGCCAUCCCAUCCCAUCACAUCCAUUC ...((((.......))--))...........(((((((.............((....)).....(((((((((.......)))))).)))..............))))))).... ( -27.90, z-score = -2.76, R) >droEre2.scaffold_4690 10314097 109 + 18748788 ----CCGAAUCCUCCG--GGUAAAUAUUUAAGGAUGUGUUACCACCUAUUUACCAUCAUUUCUCCUUGGCGGCUGCAAGUGCCGCCUAAGCCAGCCCAUCCCAUUCCAACUAGUC ----..((((.....(--(((...........((((.((............)))))).......(((((((((.......)))))).)))...)))).....))))......... ( -24.30, z-score = -1.03, R) >droPer1.super_13 772863 95 - 2293547 AUGGCAUACUCCUCUAUUGGUAAAUAUUUAAGG--GCACUAUC-UUCGAAUACCAGCACCCGUC--UGGCGGCUUAAAGUGCAGCCUAAGCCGUCGAGUG--------------- ......(((((.....((((((....((..(((--(.....))-))..))))))))........--.(((((((((..........))))))))))))))--------------- ( -22.40, z-score = 0.28, R) >dp4.chrXL_group1e 7033734 95 + 12523060 AUGGCAUACUCCUCUAUUGGUAAAUAUUUAAGG--GCACUAUC-UUCGCACACCAGCACCCGUC--UGGCGGCUUAAAGUGCAGCCUAAGCCGUCGAGUG--------------- (((((.....((......))..........(((--(((((...-...((.(.(((((....).)--))).)))....)))))..)))..)))))......--------------- ( -22.00, z-score = 0.69, R) >consensus CUCCCCGAAUCCUCCG__GGUAAAUAUUUAAGGAUGUGUUACCACCUAUUUACCAUCAUUUCUCCUUGGCGGCUGCAAGUGCCGCCUAAGCCAUCCCAUCCCAUC_CA_CC____ ..................((((((((.....((........))...)))))))).............((((((.......))))))............................. (-15.23 = -16.28 + 1.06)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:03:29 2011