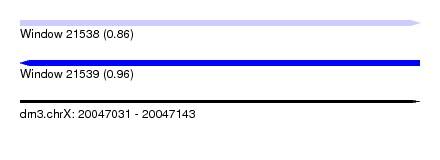
| Sequence ID | dm3.chrX |
|---|---|
| Location | 20,047,031 – 20,047,143 |
| Length | 112 |
| Max. P | 0.958968 |

| Location | 20,047,031 – 20,047,143 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.51 |
| Shannon entropy | 0.21128 |
| G+C content | 0.39817 |
| Mean single sequence MFE | -26.28 |
| Consensus MFE | -19.59 |
| Energy contribution | -20.03 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.861447 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20047031 112 + 22422827 GUUGCCUU-----UAC---UCCGGCCUCCGUUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCUG ((((((..-----...---...)))...(((((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........))))))))))...... ( -26.20, z-score = -3.02, R) >droAna3.scaffold_13417 1709168 120 + 6960332 GUUGCCAUUCGUGUGUCGGUGUGUCCUUCCGUUGCGUGCAAUUCCCACUGCAAUGCAAUAUUUAUUUGCAUUUCCAAUUAGCAACCGACUUGGAAACAUAAAUUUUAAACGACCCGAUGG ....((((.((.(.((((((((.(((.((.(((((..(((........)))(((((((.......)))))))........))))).))...))).))))..........))))))))))) ( -30.90, z-score = -1.78, R) >droEre2.scaffold_4690 10174392 112 + 18748788 GUUGCCUU-----UAC---UCCGGCCGUCGCUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCCA ...(((..-----...---...)))..(((.((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........)))).)))........ ( -22.70, z-score = -1.19, R) >droYak2.chrX 10392112 117 - 21770863 GUUGCCUUCUGUGUGU---UGCGGCCUCCGUUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCUA (((((...(.....).---.)))))...(((((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........)))))))......... ( -25.50, z-score = -1.00, R) >droSec1.super_8 2273447 112 + 3762037 GUUGCCUU-----UAC---UCCGGCCUCCGUUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCUA ((((((..-----...---...)))...(((((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........))))))))))...... ( -26.20, z-score = -3.08, R) >droSim1.chrX 15425123 112 + 17042790 GUUGCCUU-----UAC---UCCGGCCUCCGUUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCUA ((((((..-----...---...)))...(((((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........))))))))))...... ( -26.20, z-score = -3.08, R) >consensus GUUGCCUU_____UAC___UCCGGCCUCCGUUUGCGUGCAAUUCCCAUUGAAAUACAAUAUUUAUUUGCAUUUCCAAUUAGCAACUGACUUGGAAACAAUAAUUGCAAACGAACAGUCUA ......................(((...(((((((((((((.....((((.....))))......))))))((((((((((...)))).)))))).........)))))))....))).. (-19.59 = -20.03 + 0.45)

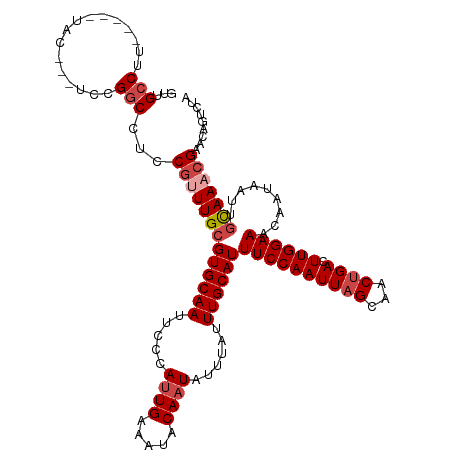
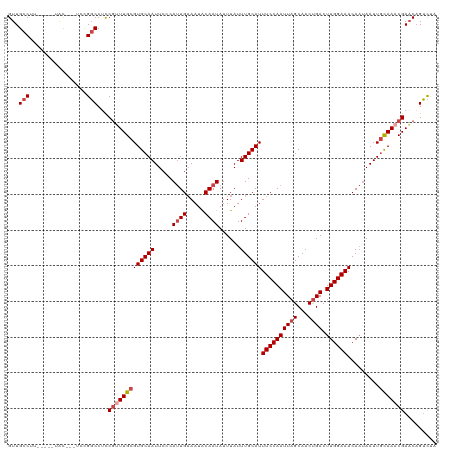
| Location | 20,047,031 – 20,047,143 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.51 |
| Shannon entropy | 0.21128 |
| G+C content | 0.39817 |
| Mean single sequence MFE | -32.19 |
| Consensus MFE | -25.30 |
| Energy contribution | -26.19 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.958968 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 20047031 112 - 22422827 CAGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAACGGAGGCCGGA---GUA-----AAGGCAAC .......(((((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).))))))))))).(((...---...-----..)))... ( -32.00, z-score = -2.71, R) >droAna3.scaffold_13417 1709168 120 - 6960332 CCAUCGGGUCGUUUAAAAUUUAUGUUUCCAAGUCGGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGCAUUGCAGUGGGAAUUGCACGCAACGGAAGGACACACCGACACACGAAUGGCAAC ((((((.((((...........(((.(((...((.(((((........(((((((.......)))))))(((((....)))))..))))).)).))).))).))))...)).)))).... ( -33.70, z-score = -1.96, R) >droEre2.scaffold_4690 10174392 112 - 18748788 UGGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAGCGACGGCCGGA---GUA-----AAGGCAAC .((.((((.(((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).)))))))))))))))...---...-----........ ( -32.60, z-score = -2.35, R) >droYak2.chrX 10392112 117 + 21770863 UAGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAACGGAGGCCGCA---ACACACAGAAGGCAAC .......(((((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).))))))))))).(((...---..........)))... ( -30.82, z-score = -1.99, R) >droSec1.super_8 2273447 112 - 3762037 UAGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAACGGAGGCCGGA---GUA-----AAGGCAAC .......(((((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).))))))))))).(((...---...-----..)))... ( -32.00, z-score = -2.86, R) >droSim1.chrX 15425123 112 - 17042790 UAGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAACGGAGGCCGGA---GUA-----AAGGCAAC .......(((((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).))))))))))).(((...---...-----..)))... ( -32.00, z-score = -2.86, R) >consensus UAGACUGUUCGUUUGCAAUUAUUGUUUCCAAGUCAGUUGCUAAUUGGAAAUGCAAAUAAAUAUUGUAUUUCAAUGGGAAUUGCACGCAAACGGAGGCCGGA___GUA_____AAGGCAAC .......(((((((((.....(((....)))(((((((.(((....(((((((((.......)))))))))..))).))))).))))))))))).(((................)))... (-25.30 = -26.19 + 0.89)

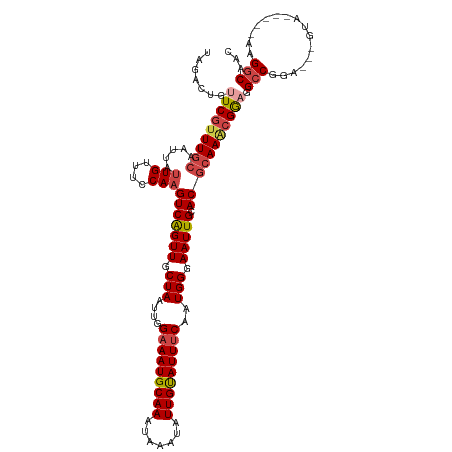
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:02:54 2011