
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,964,158 – 19,964,259 |
| Length | 101 |
| Max. P | 0.637027 |

| Location | 19,964,158 – 19,964,259 |
|---|---|
| Length | 101 |
| Sequences | 9 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 85.88 |
| Shannon entropy | 0.28909 |
| G+C content | 0.35213 |
| Mean single sequence MFE | -21.14 |
| Consensus MFE | -14.99 |
| Energy contribution | -14.88 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637027 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 19964158 101 - 22422827 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGGGCAAAUCCAACAUGCCACG--CA--CACUUAUGCACCUAUG---- .......................((((((((((((((((........)))))))))))((...((((.........))))..)--).--.....)))))......---- ( -22.10, z-score = -1.95, R) >droSim1.chrX 15353926 98 - 17042790 UUGACAUUCAUAUUACUUA-UUAUGCAUGGGAAAUU-AUUUUGAUAUGUUGAUUUCUCGCAUGGGCAAAUCCAACAUGCCACG--CA--CACUUAUGCA-CUAUG---- ...................-...(((((((((((((-(...........)))))))))((...((((.........))))..)--).--.....)))))-.....---- ( -19.10, z-score = -1.02, R) >droSec1.super_8 2190084 101 - 3762037 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGGGCAAAUCCAACAUGCCACG--CA--CACUUAUGCACCAAUG---- .......................((((((((((((((((........)))))))))))((...((((.........))))..)--).--.....)))))......---- ( -22.10, z-score = -1.75, R) >droYak2.chrX 10321942 103 + 21770863 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGGGCAAAUCCAACAUGCCACGCACA--CACUUAUGCACCUAUG---- .......................((((((((((((((((........)))))))))))((...((((.........))))..))...--.....)))))......---- ( -22.10, z-score = -1.94, R) >droEre2.scaffold_4690 10109282 101 - 18748788 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGGGCACAUCCAACAUGCCACG--CA--CACUUAUGCACCUAUG---- .......................((((((((((((((((........)))))))))))((...((((.........))))..)--).--.....)))))......---- ( -21.90, z-score = -1.92, R) >droAna3.scaffold_13417 5134016 109 + 6960332 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGAGCAAGUCCAACAUGGAGCGGGCGACUUCCAACGCAAUAAUCAAAC ((((..(((((.((((((.(((((((..(((((((((((........)))))))))))))))))).))))..)).)))))(((............))).....)))).. ( -25.70, z-score = -1.96, R) >dp4.chrXL_group1e 5267892 100 + 12523060 AUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGAGCAAAUCCAACAUGCCACACGGA--CACGGACACGAAC------- ......(((..........(((((((..(((((((((((........))))))))))))))))))....(((....((((....)).--)).)))...))).------- ( -19.60, z-score = -1.57, R) >droPer1.super_13 1293342 100 - 2293547 AUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGAGCAAAUCCAACAUGCCACACGGA--CACGGACACAGAC------- ...................(((((((..(((((((((((........))))))))))))))))))....(((....((((....)).--)).))).......------- ( -18.60, z-score = -1.20, R) >droWil1.scaffold_180777 4041844 105 + 4753960 UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGAACAAAUCCAAGAUGUGACA--UA--CAAAAAUACAAUCAUAAAAC (((....(((((((.....(((((((..(((((((((((........)))))))))))))))))).........)))))))..--..--)))................. ( -19.04, z-score = -1.60, R) >consensus UUGACAUUCAUAUUACUUAUUUAUGCAUGGGAAAUUAAUUUUGAUAUGUUGAUUUCUCGCAUGGGCAAAUCCAACAUGCCACG__CA__CACUUAUGCACCUAUG____ ...................(((((((..(((((((((((........))))))))))))))))))............................................ (-14.99 = -14.88 + -0.11)


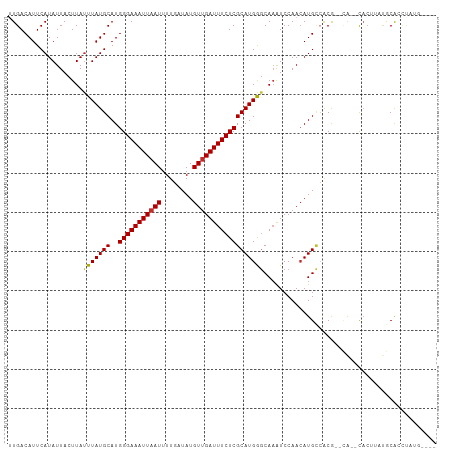
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:02:46 2011