
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,442,497 – 19,442,565 |
| Length | 68 |
| Max. P | 0.997883 |

| Location | 19,442,497 – 19,442,565 |
|---|---|
| Length | 68 |
| Sequences | 3 |
| Columns | 68 |
| Reading direction | forward |
| Mean pairwise identity | 58.42 |
| Shannon entropy | 0.57329 |
| G+C content | 0.39691 |
| Mean single sequence MFE | -19.60 |
| Consensus MFE | -17.18 |
| Energy contribution | -15.97 |
| Covariance contribution | -1.21 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.20 |
| SVM RNA-class probability | 0.997883 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
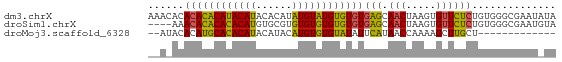
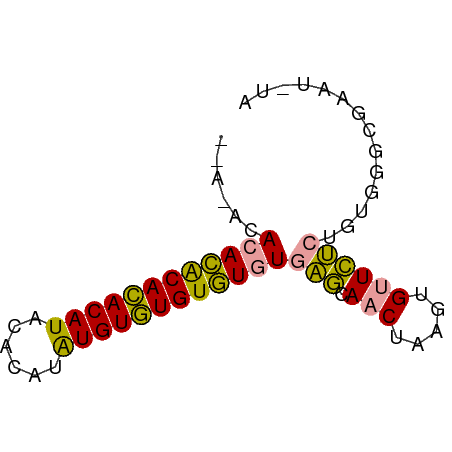
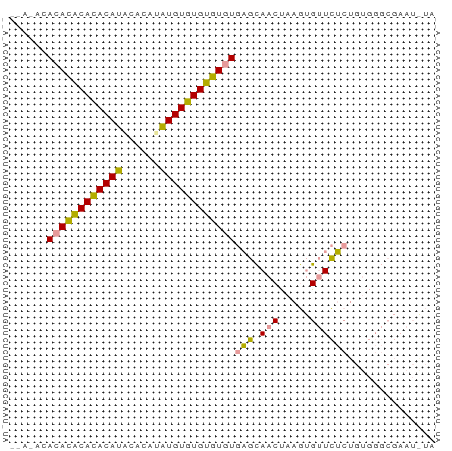
>dm3.chrX 19442497 68 + 22422827 AAACACACACACAUACAUACACAUAUGUAUGUGUGUGAGCAACUAAGUGUUCUCUGUGGGCGAAUAUA ...((((((((((((((((....)))))))))))))(((.(((.....)))))).))).......... ( -24.00, z-score = -3.24, R) >droSim1.chrX 15008244 64 + 17042790 ----AAACACACACACAUGUGCGUGUGUGUGUGUGUGAGCAACUAAGUGUUCUCUGUGGGCGAAUGUA ----..((((((((((((....))))))))))))(.(((((......))))).).............. ( -23.00, z-score = -1.47, R) >droMoj3.scaffold_6328 1541373 53 - 4453435 --AUACACAUGCACACAUACAUACAUGUGUGUAUAUUCAUAACCAAAAGCUUGCU------------- --......((((((((((......)))))))))).....................------------- ( -11.80, z-score = -1.93, R) >consensus __A_ACACACACACACAUACACAUAUGUGUGUGUGUGAGCAACUAAGUGUUCUCUGUGGGCGAAU_UA ......((((((((((((......))))))))))))(((.(((.....)))))).............. (-17.18 = -15.97 + -1.21)
| Location | 19,442,497 – 19,442,565 |
|---|---|
| Length | 68 |
| Sequences | 3 |
| Columns | 68 |
| Reading direction | reverse |
| Mean pairwise identity | 58.42 |
| Shannon entropy | 0.57329 |
| G+C content | 0.39691 |
| Mean single sequence MFE | -18.80 |
| Consensus MFE | -17.96 |
| Energy contribution | -16.97 |
| Covariance contribution | -0.99 |
| Combinations/Pair | 1.37 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.96 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.66 |
| SVM RNA-class probability | 0.993978 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
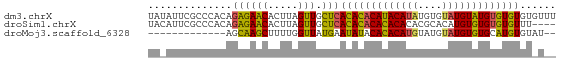
>dm3.chrX 19442497 68 - 22422827 UAUAUUCGCCCACAGAGAACACUUAGUUGCUCACACACAUACAUAUGUGUAUGUAUGUGUGUGUGUUU ..........(((.((((((.....))).)))(((((((((((((....))))))))))))))))... ( -23.80, z-score = -3.08, R) >droSim1.chrX 15008244 64 - 17042790 UACAUUCGCCCACAGAGAACACUUAGUUGCUCACACACACACACACGCACAUGUGUGUGUGUUU---- ..............((((((.....))).)))..(((((((((((......)))))))))))..---- ( -20.10, z-score = -2.72, R) >droMoj3.scaffold_6328 1541373 53 + 4453435 -------------AGCAAGCUUUUGGUUAUGAAUAUACACACAUGUAUGUAUGUGUGCAUGUGUAU-- -------------.........................((((((((((......))))))))))..-- ( -12.50, z-score = 0.15, R) >consensus UA_AUUCGCCCACAGAGAACACUUAGUUGCUCACACACACACAUAUGUGUAUGUGUGUGUGUGU_U__ ..............((((((.....))).)))(((((((((((((....)))))))))))))...... (-17.96 = -16.97 + -0.99)
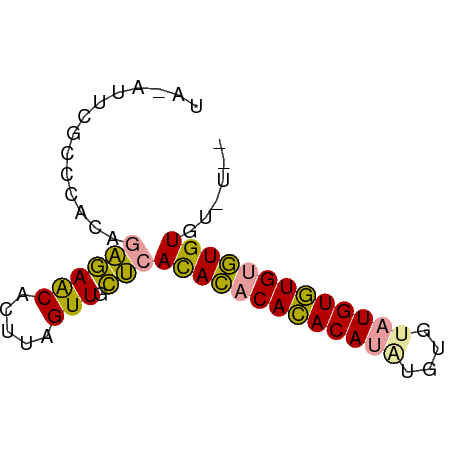
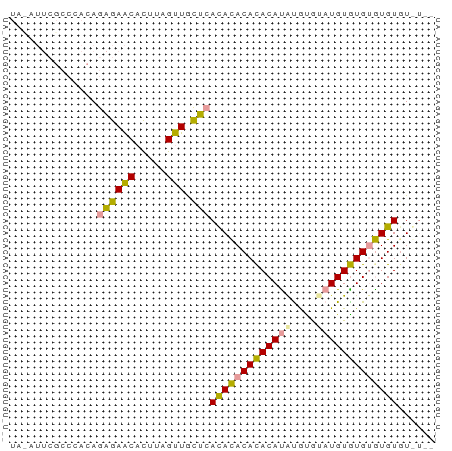
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:01:19 2011