
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,441,642 – 19,441,750 |
| Length | 108 |
| Max. P | 0.507785 |

| Location | 19,441,642 – 19,441,750 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 66.19 |
| Shannon entropy | 0.60272 |
| G+C content | 0.39087 |
| Mean single sequence MFE | -18.46 |
| Consensus MFE | -7.64 |
| Energy contribution | -8.08 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.507785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 19441642 108 + 22422827 CAAU-GCCAGGACGAAUAUGCCAGAAAAUGGGGUAUAACUGGAUUU-----GGAAAGAAUUAAGGGAACGAUUGAGAUAGUAACUUACAUAAGUAGCCGACUAAAGCAGACGAC-- (((.-.((((.....((((.(((.....))).))))..))))..))-----)................((.(((...((((...((((....))))...))))...))).))..-- ( -20.00, z-score = -1.80, R) >droEre2.scaffold_4690 9678805 81 + 18748788 CAAU-GCCAGGAGUAAUAUGCCAGAAAAAGGGGUACAACUGAA--U-----AGAUA------------CGAUUAGGAUGGUAACAUAAAUGGGAACAUAAA--------------- ...(-((((..(((..(((.((.......)).)))..)))...--.-----.....------------(.....)..)))))......(((....)))...--------------- ( -10.90, z-score = -0.52, R) >droYak2.chrX 9875352 112 - 21770863 CAAUUGCCAGGACGAAAAUGCCUGAAAAAGGCAUAUAAGUGGAUUUUGGCCAGGAAAUUACCAACUUACGAAAUUAGUAAUAAGUAACAUAAAUAGACGACUGAAGCAGGCC---- .....(((....((...((((((.....)))))).(((((.((((((......))))))....)))))))...(((((.....((...........)).)))))....))).---- ( -19.70, z-score = -0.73, R) >droSec1.super_8 1723931 109 + 3762037 CAAU-GCCAGGACGAAUAUGCCAGAAAAUGGGGUAUAACUGGAUUU-----GGAUAGAAACU-GAGUACGAUUAAGAUAGUAACUUACAUAAGUAGACGACUCUAAGGCAGAACAC ...(-(((.......((((.(((.....))).))))...((((.((-----(..((.....(-((((((..........)).)))))......))..))).)))).))))...... ( -19.70, z-score = -1.24, R) >droSim1.chrX 15007315 108 + 17042790 CAAU-GCCAGGACGAAUACGCCAGAAAAUGGGGUAUAACUGGAUUU-----GGAUAGAAACU-GAGUUCGAUUAAGAUAGUAACUUACAUAAGUAGACGACUCUAAGGCA-AACAC ...(-(((.((.((....))))......((((((.(((.((((((.-----.(.......).-.)))))).))).....((.((((....))))..)).)))))).))))-..... ( -22.00, z-score = -1.79, R) >consensus CAAU_GCCAGGACGAAUAUGCCAGAAAAUGGGGUAUAACUGGAUUU_____GGAUAGAAACU_GAGUACGAUUAAGAUAGUAACUUACAUAAGUAGACGACUCAAACACA_AAC__ ......((((.....((((.(((.....))).))))..)))).......................................................................... ( -7.64 = -8.08 + 0.44)

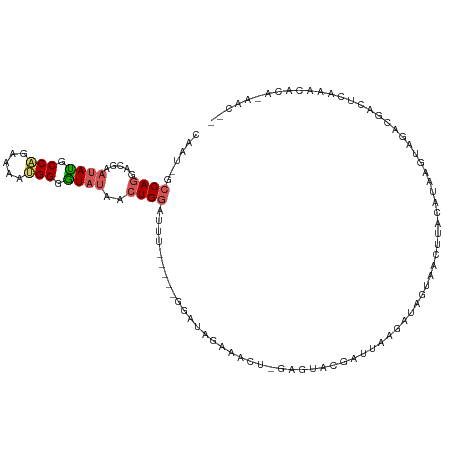

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:01:17 2011