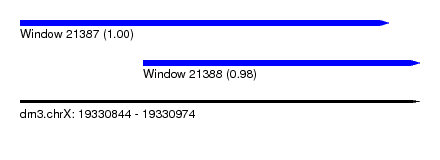

| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,330,844 – 19,330,974 |
| Length | 130 |
| Max. P | 0.995231 |

| Location | 19,330,844 – 19,330,964 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.06 |
| Shannon entropy | 0.28723 |
| G+C content | 0.53355 |
| Mean single sequence MFE | -46.90 |
| Consensus MFE | -38.75 |
| Energy contribution | -39.87 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.84 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.78 |
| SVM RNA-class probability | 0.995231 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
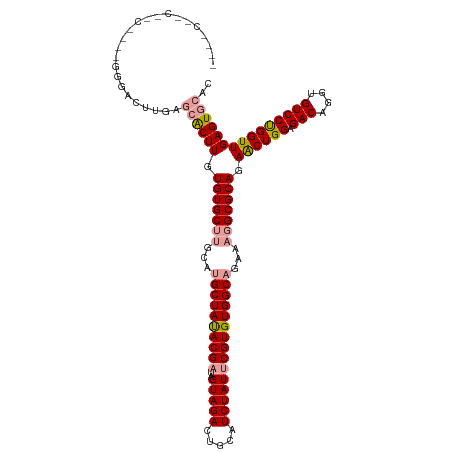
>dm3.chrX 19330844 120 + 22422827 CCUUCUCCUGCACUUGGGACUUGAGCGCUUGUGUGCUUGCUUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCCGGUUGAGUGCAC ........((((((((((((........((.(((((((..(((((((((((..(((((.....))))))))))))))))..))))))).))(((....)))..)))))....))))))). ( -51.20, z-score = -3.53, R) >droEre2.scaffold_4690 9579553 104 + 18748788 ---------------GGCACAUGUGCACUUGUGUGCAUGCAGGCUACACGACAGUAGACUGCAUCUAUCCGUGUGGCAGAA-GGCGCAGAGCUGGAGACAGGUGUCCUGGUUGAGUGCAC ---------------(.(((((((((((....)))))(((((.((((......)))).)))))......)))))).)....-.((((..(((..(.(((....))))..)))..)))).. ( -42.40, z-score = -1.16, R) >droSim1.chrX 14955899 116 + 17042790 ----CCUCAUCCUACUGGACUUGGAAACUUGUGUGCUUGCAUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCUGGUUGAGUGCAC ----........((((.((((.(((.((((.(((((((...((((((((((..(((((.....)))))))))))))))...))))))).))(((....))))).))).)))).))))... ( -47.10, z-score = -3.82, R) >consensus ____C__C__C____GGGACUUGAGCACUUGUGUGCUUGCAUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCUGGUUGAGUGCAC ........................((((((.(((((((...((((((((((..(((((.....)))))))))))))))...))))))).((((((.(((....))))))))))))))).. (-38.75 = -39.87 + 1.12)


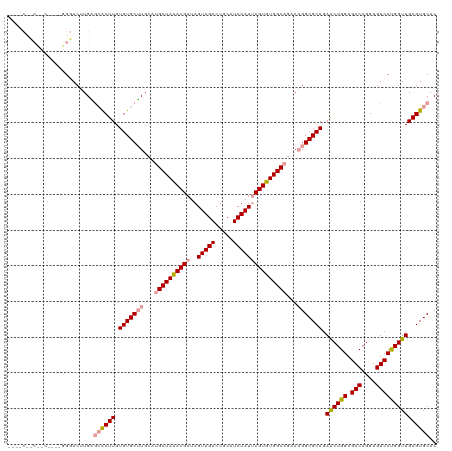
| Location | 19,330,884 – 19,330,974 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 87.94 |
| Shannon entropy | 0.16261 |
| G+C content | 0.53022 |
| Mean single sequence MFE | -32.30 |
| Consensus MFE | -31.53 |
| Energy contribution | -31.20 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.98 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.976041 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 19330884 90 + 22422827 UUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCCGGUUGAGUGCACACCCAUAUCC------ (((((((((((..(((((.....))))))))))))))))....((((..((((((.(((....)))))))))..))))............------ ( -34.60, z-score = -3.33, R) >droEre2.scaffold_4690 9579578 88 + 18748788 AGGCUACACGACAGUAGACUGCAUCUAUCCGUGUGGCAGAA-GGCGCAGAGCUGGAGACAGGUGUCCUGGUUGAGUGCACACCCACACC------- ..((((((((...(((((.....))))).))))))))....-.((((..(((..(.(((....))))..)))..))))...........------- ( -30.60, z-score = -0.48, R) >droSim1.chrX 14955935 96 + 17042790 AUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCUGGUUGAGUGCACACCCAUACCCAUAUCC .((((((((((..(((((.....))))))))))))))).....((((..(((..(.(((....))))..)))..)))).................. ( -31.70, z-score = -1.80, R) >consensus AUGCUAUACGAUAGUAGACUGCAUCUAUUCGUGUGGCAGAAAGGCGCAGAACUGGAGACAGGUGUCCUGGUUGAGUGCACACCCAUACCC______ ..(((((((((..(((((.....))))))))))))))......((((..((((((.(((....)))))))))..)))).................. (-31.53 = -31.20 + -0.33)

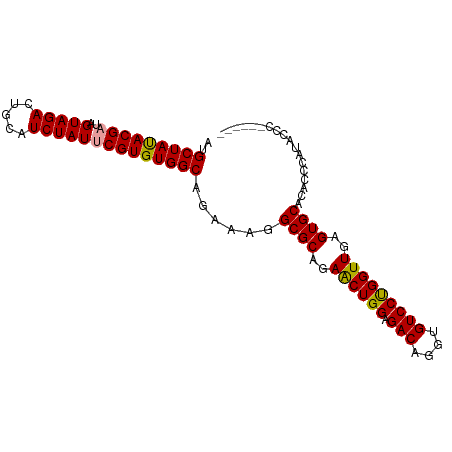
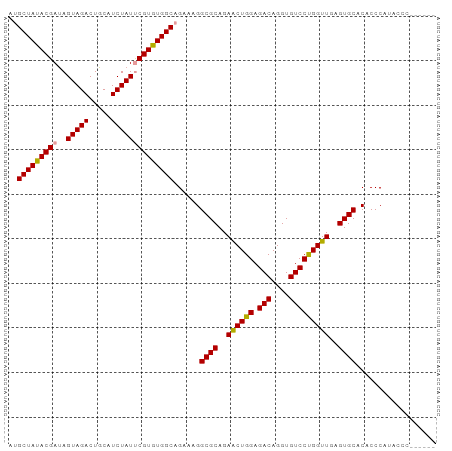
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:00:51 2011