
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,271,948 – 19,272,053 |
| Length | 105 |
| Max. P | 0.796058 |

| Location | 19,271,948 – 19,272,053 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.45 |
| Shannon entropy | 0.19591 |
| G+C content | 0.57338 |
| Mean single sequence MFE | -41.32 |
| Consensus MFE | -35.68 |
| Energy contribution | -35.96 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.86 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.796058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
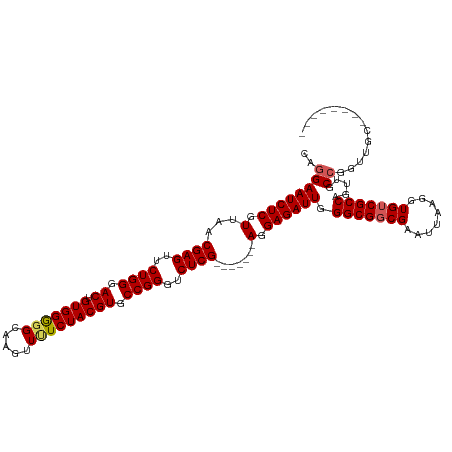
>dm3.chrX 19271948 105 + 22422827 CAGGAAUCUCGUUAACGAGUUCUGGGACUGUGGGUGCAAGUUUUCUACGUGCCGGGUCUCG-------AGGAGAUUGGGCGGCGAAUUAAGGUGUCGCCAGUUGCCGGUUGC-------- ..((((((((.(...((((..((((.((.(((((.........))))))).))))..))))-------).)))))).(((((((........))))))).....))......-------- ( -37.10, z-score = -1.31, R) >droEre2.scaffold_4690 9523357 113 + 18748788 CAGGAAUCUCGUUAACGAGUUCUGGGACUGUGGGGGCAAGUUUUCUACGUGCCGGGUCUCG-------AGGAGAUUGGGCGGCGAGUUAAGGUGUCGCCAGUAGCCGGUUUCGGGUUUCG ..(((((((((....))))..(((..(((((((.((((..((..((.((((((.((((((.-------..)))))).))).)))))..))..)))).))).....))))..)))))))). ( -42.90, z-score = -2.02, R) >droYak2.chrX 9713503 120 - 21770863 CAGGAAUCUCGUUAACGAGUUCUGGGACUGUGGAGGCAAGUUUUCUACGUGCCGGGUCUCGGCUCUUGACGAGAUUGGGCGCCGAAUUAAGGUGUCGCCAGUAGCCGAUUGCCGAUUGCC ..(((((((((((((((((..((((.((.(((((((.....))))))))).))))..))))....))))))))))).((((((.......)))))).)).(((..((.....))..))). ( -46.10, z-score = -1.70, R) >droSec1.super_8 1564313 105 + 3762037 CACGAAUCUCGUUAACGAGUUCUGGGACUGUGGGGGCAAGUUCUCUACGUGCCGGGUCUCG-------AGGAGAUUGGGCGGCGAAUUAAGGUGUCGCCAGUUGCCGGUUGC-------- ..(.((((((.(...((((..((((.((.(((((((.....))))))))).))))..))))-------).)))))).).((((((.....((.....))..)))))).....-------- ( -39.90, z-score = -1.56, R) >droSim1.chrX_random 5024481 105 + 5698898 CAGGAAUCUCGUUAACGAGUUCUGGGACUGUGGGGGCAAGUUCUCUACGUGCCGGGUCUCG-------AGGAGAUUGGGCGGCGAAUUAAGGUGUCGCCAGUUGCCGGUGCC-------- ..((((((((.(...((((..((((.((.(((((((.....))))))))).))))..))))-------).)))))).(((((((........))))))).....))......-------- ( -40.60, z-score = -1.53, R) >consensus CAGGAAUCUCGUUAACGAGUUCUGGGACUGUGGGGGCAAGUUUUCUACGUGCCGGGUCUCG_______AGGAGAUUGGGCGGCGAAUUAAGGUGUCGCCAGUUGCCGGUUGC________ ..((((((((.(...((((..((((.((.(((((((.....))))))))).))))..)))).......).)))))).(((((((........))))))).....)).............. (-35.68 = -35.96 + 0.28)


| Location | 19,271,948 – 19,272,053 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.45 |
| Shannon entropy | 0.19591 |
| G+C content | 0.57338 |
| Mean single sequence MFE | -28.14 |
| Consensus MFE | -23.10 |
| Energy contribution | -23.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.741269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
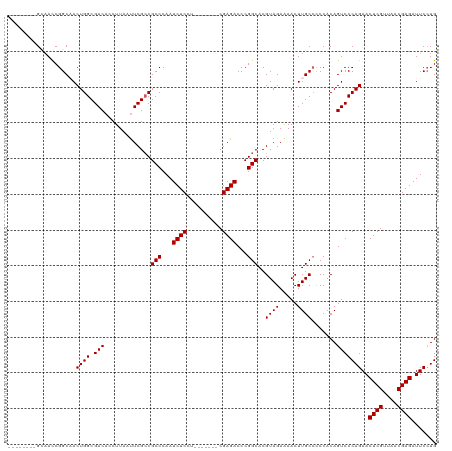
>dm3.chrX 19271948 105 - 22422827 --------GCAACCGGCAACUGGCGACACCUUAAUUCGCCGCCCAAUCUCCU-------CGAGACCCGGCACGUAGAAAACUUGCACCCACAGUCCCAGAACUCGUUAACGAGAUUCCUG --------((.....))..((((.(((..........((((.....((((..-------.))))..))))..((((.....)))).......)))))))..((((....))))....... ( -25.10, z-score = -1.92, R) >droEre2.scaffold_4690 9523357 113 - 18748788 CGAAACCCGAAACCGGCUACUGGCGACACCUUAACUCGCCGCCCAAUCUCCU-------CGAGACCCGGCACGUAGAAAACUUGCCCCCACAGUCCCAGAACUCGUUAACGAGAUUCCUG ......(((....)))((((.(((((.........)))))(((...((((..-------.))))...)))..))))......................(((((((....)))).)))... ( -26.20, z-score = -1.94, R) >droYak2.chrX 9713503 120 + 21770863 GGCAAUCGGCAAUCGGCUACUGGCGACACCUUAAUUCGGCGCCCAAUCUCGUCAAGAGCCGAGACCCGGCACGUAGAAAACUUGCCUCCACAGUCCCAGAACUCGUUAACGAGAUUCCUG (((....((...((((((..(((((((.((.......)).).......))))))..))))))...))((((.((.....)).))))......)))...(((((((....)))).)))... ( -35.01, z-score = -1.45, R) >droSec1.super_8 1564313 105 - 3762037 --------GCAACCGGCAACUGGCGACACCUUAAUUCGCCGCCCAAUCUCCU-------CGAGACCCGGCACGUAGAGAACUUGCCCCCACAGUCCCAGAACUCGUUAACGAGAUUCGUG --------((....(((((..(((((.........)))))(((...((((..-------.))))...)))...........)))))............(((((((....)))).))))). ( -27.40, z-score = -1.82, R) >droSim1.chrX_random 5024481 105 - 5698898 --------GGCACCGGCAACUGGCGACACCUUAAUUCGCCGCCCAAUCUCCU-------CGAGACCCGGCACGUAGAGAACUUGCCCCCACAGUCCCAGAACUCGUUAACGAGAUUCCUG --------((....(((((..(((((.........)))))(((...((((..-------.))))...)))...........))))).)).........(((((((....)))).)))... ( -27.00, z-score = -1.48, R) >consensus ________GCAACCGGCAACUGGCGACACCUUAAUUCGCCGCCCAAUCUCCU_______CGAGACCCGGCACGUAGAAAACUUGCCCCCACAGUCCCAGAACUCGUUAACGAGAUUCCUG ...................((((.(((.............(((...((((..........))))...)))..((((.....)))).......)))))))..((((....))))....... (-23.10 = -23.10 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:00:18 2011