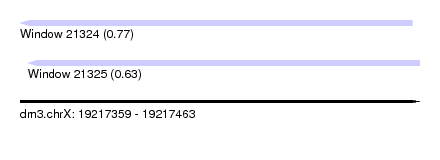

| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,217,359 – 19,217,463 |
| Length | 104 |
| Max. P | 0.765854 |

| Location | 19,217,359 – 19,217,461 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 77.06 |
| Shannon entropy | 0.39794 |
| G+C content | 0.37080 |
| Mean single sequence MFE | -25.58 |
| Consensus MFE | -15.76 |
| Energy contribution | -16.16 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.765854 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
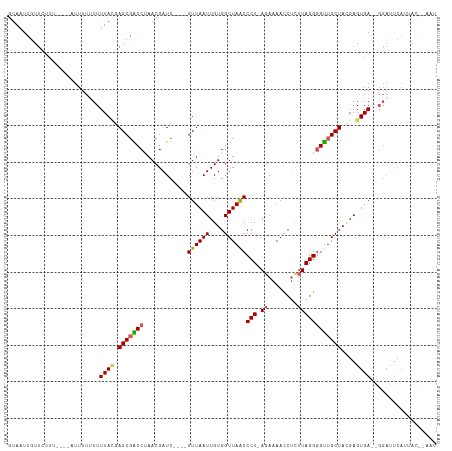
>dm3.chrX 19217359 102 - 22422827 GUAAUUGUUC--------UUUUUUUUUACGAGCGACCAAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAACCUCUUAGGGGUUGCUACGAGUGA--GGACUCAAUACUUAAU (((...((((--------((.......((((.((......)).))----))...(((((((...((((-(((.....)).)))))...)))))))..))--))))....)))..... ( -25.20, z-score = -1.60, R) >droEre2.scaffold_4690 9474233 104 - 18748788 GUAAUUGUUUUU-----AUUUUGUUUUAAGAGCGGCCCAACGAUCGUGGGUUAAUUGUGGUUAACCCU-AGAAAACCAUUUAGGGGCUGCUAUGAGUGA--GGAUUCAGGAU----- ......((((((-----((((.........((((((((.((....))((((((((....)))))))).-..............))))))))..))))))--)))).......----- ( -28.70, z-score = -1.61, R) >droYak2.chrX 9662989 92 + 21770863 GUAAUAGUAAUUU---AAUUUAAUUUUAUGAGCCGCCUAACGAUC----GUUAAUUGUGGUUAGCCCUAAAAAAACCUCAUAGGGGUUGCUAGGAGUGA------------------ ((((...(((...---...)))...)))).((((((.((((....----))))...)))))).((((((....(((((.....)))))..)))).))..------------------ ( -18.90, z-score = -0.43, R) >droSec1.super_8 1510807 110 - 3762037 GUAAUUGUUCUUUUAUUUUUUUUUUUUACGAGCGACCUAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAAUCUCUUAGGGGUUGCUACGAGUGA--GGAUUCAUUACCAAAU (((((.((((((..................(((((.(....).))----)))..(((((((...((((-(((.....)).)))))...)))))))..))--))))..)))))..... ( -26.90, z-score = -2.18, R) >droSim1.chrX 14857067 110 - 17042790 GUAAUUGUUCUUUUA--UUUUUUUUUUACGAGCGACCUAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAACCUCUUAGGUGUUGCUACGAGUGAGGGGAUUCAUUACCUAAU (((((.(((((((((--(((..........(((((.(....).))----)))....(((((.((((((-(((.....)).)))).))))))))))))))))))))..)))))..... ( -28.20, z-score = -2.27, R) >consensus GUAAUUGUUCUUU____AUUUUUUUUUACGAGCGACCUAACGAUC____GUUAAUUGUGGUUAACCCU_AGAAAACCUCUUAGGGGUUGCUACGAGUGA__GGAUUCAUUAC__AAU .........................((((.(((((((............((((((....))))))(((.(((.....))).))))))))))....)))).................. (-15.76 = -16.16 + 0.40)


| Location | 19,217,361 – 19,217,463 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 78.18 |
| Shannon entropy | 0.38134 |
| G+C content | 0.37736 |
| Mean single sequence MFE | -26.34 |
| Consensus MFE | -15.76 |
| Energy contribution | -16.16 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.630886 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
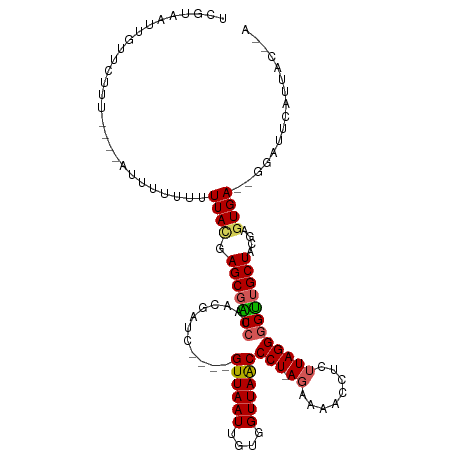
>dm3.chrX 19217361 102 - 22422827 UCGUAAUUGUUC--------UUUUUUUUUACGAGCGACCAAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAACCUCUUAGGGGUUGCUACGAGUGA--GGACUCAAUACUUA ..(((...((((--------((.......((((.((......)).))----))...(((((((...((((-(((.....)).)))))...)))))))..))--))))....)))... ( -25.50, z-score = -1.37, R) >droEre2.scaffold_4690 9474233 106 - 18748788 UCGUAAUUGUUUUU-----AUUUUGUUUUAAGAGCGGCCCAACGAUCGUGGGUUAAUUGUGGUUAACCCU-AGAAAACCAUUUAGGGGCUGCUAUGAGUGA--GGAUUCAGGAU--- ((((..((((((((-----(........)))))))))....))))((.((((((..(..((((...((((-(((......)))))))...))))..)....--.)))))).)).--- ( -29.00, z-score = -1.42, R) >droYak2.chrX 9662989 94 + 21770863 UCGUAAUAGUAAUUU---AAUUUAAUUUUAUGAGCCGCCUAACGAUC----GUUAAUUGUGGUUAGCCCUAAAAAAACCUCAUAGGGGUUGCUAGGAGUGA---------------- ((((((...(((...---...)))...))))))(((((.((((....----))))...)))))..((((((....(((((.....)))))..)))).))..---------------- ( -21.50, z-score = -1.12, R) >droSec1.super_8 1510809 110 - 3762037 UCGUAAUUGUUCUUUUAUUUUUUUUUUUUACGAGCGACCUAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAAUCUCUUAGGGGUUGCUACGAGUGA--GGAUUCAUUACCAA ..(((((.((((((..................(((((.(....).))----)))..(((((((...((((-(((.....)).)))))...)))))))..))--))))..)))))... ( -27.20, z-score = -2.04, R) >droSim1.chrX 14857069 110 - 17042790 UCGUAAUUGUUCUUUUA--UUUUUUUUUUACGAGCGACCUAACGAUC----GUUAAUUGUGGUUAACCCU-AGAAAACCUCUUAGGUGUUGCUACGAGUGAGGGGAUUCAUUACCUA ..(((((.(((((((((--(((..........(((((.(....).))----)))....(((((.((((((-(((.....)).)))).))))))))))))))))))))..)))))... ( -28.50, z-score = -2.13, R) >consensus UCGUAAUUGUUCUUU____AUUUUUUUUUACGAGCGACCUAACGAUC____GUUAAUUGUGGUUAACCCU_AGAAAACCUCUUAGGGGUUGCUACGAGUGA__GGAUUCAUUAC__A ...........................((((.(((((((............((((((....))))))(((.(((.....))).))))))))))....))))................ (-15.76 = -16.16 + 0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 02:00:01 2011