
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,197,662 – 19,197,712 |
| Length | 50 |
| Max. P | 0.544801 |

| Location | 19,197,662 – 19,197,712 |
|---|---|
| Length | 50 |
| Sequences | 12 |
| Columns | 50 |
| Reading direction | reverse |
| Mean pairwise identity | 78.58 |
| Shannon entropy | 0.48731 |
| G+C content | 0.62000 |
| Mean single sequence MFE | -17.22 |
| Consensus MFE | -11.80 |
| Energy contribution | -11.67 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.62 |
| Mean z-score | -0.68 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.544801 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
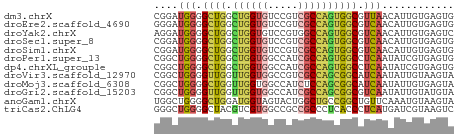
>dm3.chrX 19197662 50 - 22422827 CGGAUGGGGCUGGCUGGUGUCCGUCGCCAGUGGCGUUAACAUUGUGAGUG ....(((.((((.((((((.....)))))))))).)))............ ( -14.30, z-score = 0.10, R) >droEre2.scaffold_4690 9455251 50 - 18748788 GGGAUGGGGCUGGCUGGUGUCCGUCGCCAGUGGCGUCAACAUUGUGAGUG ....(((.((((.((((((.....)))))))))).)))............ ( -16.40, z-score = -0.28, R) >droYak2.chrX 9642465 50 + 21770863 AGGAUGGGGCUGGCUGGUGUCCGUGGCCAGUGGCGUCAACAUUGUGAGUC ..((((..(((((((.........)))))))..))))............. ( -15.60, z-score = -0.34, R) >droSec1.super_8 1491378 50 - 3762037 CGGAUGGGGCUGGCUGGUGUCCGUCGCCAGUGGCGUCAACAUUGUGAGUG ....(((.((((.((((((.....)))))))))).)))............ ( -16.40, z-score = -0.30, R) >droSim1.chrX 14837336 50 - 17042790 CGGAUGGGGCUGGCUGGUGUCCGUCGCCAGUGGCGUCAACAUUGUGAGUG ....(((.((((.((((((.....)))))))))).)))............ ( -16.40, z-score = -0.30, R) >droPer1.super_13 2085497 50 + 2293547 CGGCUGGGGCUGGCUGGUGGCCAUCGCCAGUGGCCUCAAUAUCGUGAGUG ....((((((((.((((((.....))))))))))))))....(....).. ( -20.60, z-score = -1.09, R) >dp4.chrXL_group1e 8356613 50 - 12523060 CGGCUGGGGCUGGCUGGUGGCCAUCGCCAGUGGCCUCAAUAUCGUGAGUG ....((((((((.((((((.....))))))))))))))....(....).. ( -20.60, z-score = -1.09, R) >droVir3.scaffold_12970 9408214 50 - 11907090 CGGCUGGGGUUGGUUGGUGGCCGUCGCCAGCGGCAUCAAUAUUGUAAGUA ..(((((.(.((((.....)))).).))))).(((.......)))..... ( -15.80, z-score = -0.70, R) >droMoj3.scaffold_6308 2036881 50 + 3356042 CGGCUGGGGCUGGUUGGUGGCCAUCUCCAGCGGCAUCAAUAUUGUGAGUA ..(((((((.((((.....)))).))))))).((.(((......))))). ( -18.80, z-score = -1.43, R) >droGri2.scaffold_15203 7000517 50 + 11997470 CGGCUGGGGUUGGUUGGUGGCCAUCGCCAGCGGCGUCAAUAUUGUAUGUA ..(((((.(.((((.....)))).).))))).((((((....)).)))). ( -17.50, z-score = -1.11, R) >anoGam1.chrX 9087865 50 + 22145176 UGGCUGGGGCUGGAUGGUAGUACUGGCUGCCGGCUGUUCAAAUGUAAGUA .(((((((((..(((....)).)..))).))))))............... ( -15.30, z-score = -0.75, R) >triCas2.ChLG4 11441206 50 - 13894384 GGGCUGGGGCUACGUCGUGGCCGCCGCCCUCACCCUCAUGAUCGUAAGUC ((((.((((((((...)))))).))))))..................... ( -18.90, z-score = -0.84, R) >consensus CGGCUGGGGCUGGCUGGUGGCCGUCGCCAGUGGCGUCAACAUUGUGAGUG ....(((.((((.((((((.....)))))))))).)))............ (-11.80 = -11.67 + -0.12)

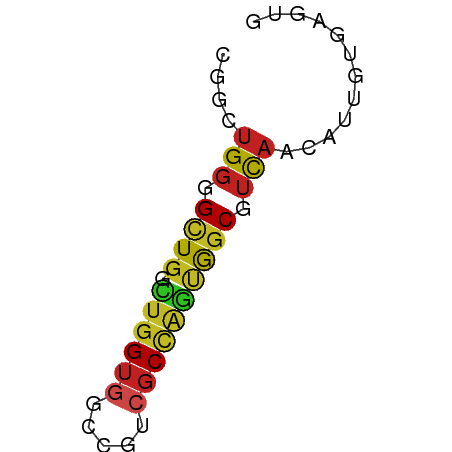

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:59:55 2011