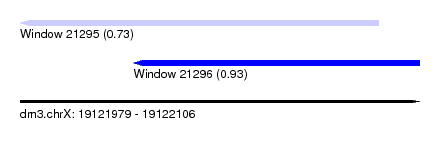
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,121,979 – 19,122,106 |
| Length | 127 |
| Max. P | 0.927809 |

| Location | 19,121,979 – 19,122,093 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 92.65 |
| Shannon entropy | 0.12687 |
| G+C content | 0.52000 |
| Mean single sequence MFE | -39.86 |
| Consensus MFE | -33.60 |
| Energy contribution | -33.16 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.730202 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
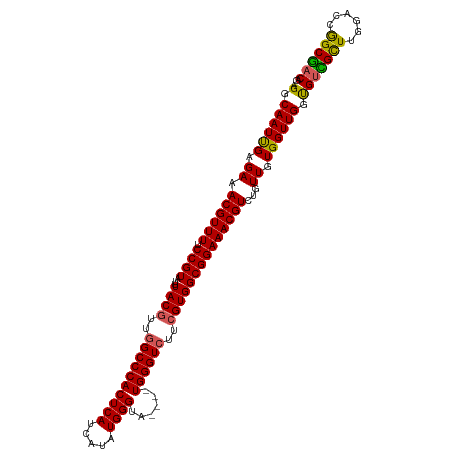
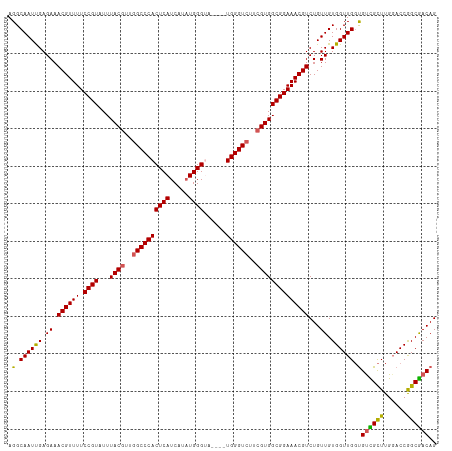
>dm3.chrX 19121979 114 - 22422827 AGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGUAUGGGUGGGUAUUCGUGGCGGAAACGUCUGUUGUGGUUGGUGUCGCUUGUACCAGCGACAG ..(((((..(((..(((((.((((...((((...(((((((((((......).))))))))))...)))))))))))))))))))))......(((((((......))))))). ( -43.20, z-score = -3.03, R) >droSim1.chrX 14787245 110 - 17042790 AGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGCA----UGGGUCUUCGUGGCGGAAACGUCUGUUGUGGUUGGUGUCGCUUGGACCGGCGACAG ..(((((..(((..(((((.((((...((((..(((((((((((...)))))..----))))))..)))))))))))))))))))))......(((((((......))))))). ( -38.70, z-score = -1.45, R) >droSec1.super_8 1415725 110 - 3762037 AGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGCA----UGGGUCUUCGUGGCGGAAACGUCUGUUGUGGUUGGUGUCGCUUGGACCGGCGACAG ..(((((..(((..(((((.((((...((((..(((((((((((...)))))..----))))))..)))))))))))))))))))))......(((((((......))))))). ( -38.70, z-score = -1.45, R) >droYak2.chrX 9567336 110 + 21770863 AGGCAAUCGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUUUGGGUA----UGGGUCGCCGUGGCGGAAACGUCUGUUGUGGUUGCUGUUGUUUGGACCGGCGACAG .((((((((.((.((((((.((((...((((.(((((((((((.....))))).----.)))))).))))))))))))))...)).))))))))((((((......)))))).. ( -42.00, z-score = -2.32, R) >droEre2.scaffold_4690 9382843 110 - 18748788 AGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGUA----UGGGUCGUAGUGGCGGAAACGUCUGUUGUGGUUGGCGCUGCUUGGACCGGCAACAG .............((((((.((((..(((((...((((((((((...)))))).----.)))))))))..))))))))))((((((((((((((...)))..)))).))))))) ( -36.70, z-score = -1.02, R) >consensus AGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGUA____UGGGUCUUCGUGGCGGAAACGUCUGUUGUGGUUGGUGUCGCUUGGACCGGCGACAG .(.((((((.((.((((((.((((...((((..((((((((((.....))))......))))))..))))))))))))))...)).)))))).)((((((......)))))).. (-33.60 = -33.16 + -0.44)

| Location | 19,122,015 – 19,122,106 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 95.13 |
| Shannon entropy | 0.07547 |
| G+C content | 0.46596 |
| Mean single sequence MFE | -28.78 |
| Consensus MFE | -23.52 |
| Energy contribution | -24.02 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927809 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
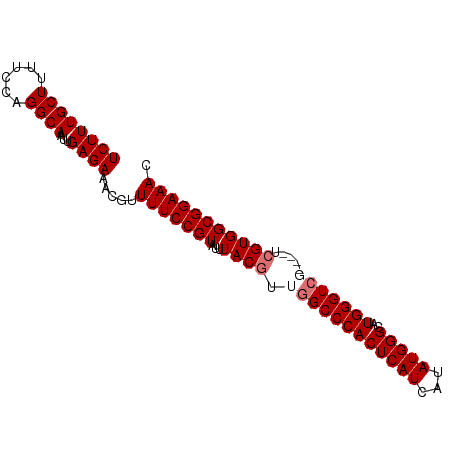
>dm3.chrX 19122015 91 - 22422827 UCUUUGCUUUUCCAGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGUAUGGGUGGGUAUUCGUGGCGGAAAC ((((((((......))))...)))).....(((((((...((((...(((((((((((......).))))))))))...))))))))))). ( -32.60, z-score = -3.48, R) >droSim1.chrX 14787281 87 - 17042790 UCUUUGCUUUUCCAGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGCAUGGGUCU----UCGUGGCGGAAAC ((((((((......))))...)))).....(((((((...((((..(((((((((((...)))))..)))))).----.))))))))))). ( -28.10, z-score = -2.56, R) >droSec1.super_8 1415761 87 - 3762037 UCUUUGCUUUUCCAGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGCAUGGGUCU----UCGUGGCGGAAAC ((((((((......))))...)))).....(((((((...((((..(((((((((((...)))))..)))))).----.))))))))))). ( -28.10, z-score = -2.56, R) >droEre2.scaffold_4690 9382879 87 - 18748788 UCUUUGCUUUUCCAGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGUAUGGGUCG----UAGUGGCGGAAAC ((((((((......))))...)))).....(((((((..(((((...((((((((((...))))))..))))))----)))..))))))). ( -26.30, z-score = -1.98, R) >consensus UCUUUGCUUUUCCAGGCAAUUGAGAAACGUUUUCCGUAUUUACGUUGGCCCACUCAUCAUAUGGGCAUGGGUCG____UCGUGGCGGAAAC ((((((((......))))...)))).....(((((((...((((..(((((((((((...)))))..))))))......))))))))))). (-23.52 = -24.02 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:59:37 2011