
| Sequence ID | dm3.chrX |
|---|---|
| Location | 19,089,585 – 19,089,680 |
| Length | 95 |
| Max. P | 0.684760 |

| Location | 19,089,585 – 19,089,680 |
|---|---|
| Length | 95 |
| Sequences | 7 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 58.78 |
| Shannon entropy | 0.71160 |
| G+C content | 0.46961 |
| Mean single sequence MFE | -27.17 |
| Consensus MFE | -13.73 |
| Energy contribution | -12.41 |
| Covariance contribution | -1.32 |
| Combinations/Pair | 1.55 |
| Mean z-score | -0.46 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.684760 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 19089585 95 - 22422827 ----AGACGUGAGUGUGUAAGUGGGUAUGGGUAUGCUUAACAAUGCCUAACAUAAACACCUGAACAAAGGUGUGCGGC-GUCUCCUUGUGCGCGUGCAAA---------- ----....((..(((..((((.(((..(((((((........)))))))......((((((......)))))).....-..)))))))..)))..))...---------- ( -29.60, z-score = -0.87, R) >droSim1.chrX 14752226 87 - 17042790 ----AUGCGUGAGUGUGUAAGUGGGUAUGGGUAUGCUUAACAAUGCCUA--------ACCUGGACAAAGGUGUGCGGC-AUCUCCGUGUGCGCGUGCAAA---------- ----((((....))))........(((((.((((((......(((((..--------((((......))))....)))-))....)))))).)))))...---------- ( -26.90, z-score = 0.05, R) >droSec1.super_8 1384022 87 - 3762037 ----AUGCGUGAGUGUGUAAGUGGGUAUGGGUAUGCUUAACAAUGCCUA--------ACCUGGACAAAGGUGUGCGGC-AUCUCCGUGUGCGCGUGCAAA---------- ----((((....))))........(((((.((((((......(((((..--------((((......))))....)))-))....)))))).)))))...---------- ( -26.90, z-score = 0.05, R) >droYak2.chrX 17678765 88 - 21770863 ------AUGUGUGAGUGUGAGUGUGUUUGGGUAUG-----CAAUGCUUAACAUAAACACCUGAUUAAAGGUGUGCGGC-AUCUCCGUGUGCGCGUGCCGU---------- ------.(((((((((((..(((((......))))-----).)))))).))))).((((((......))))))(((((-((...((....)).)))))))---------- ( -28.10, z-score = -0.98, R) >droEre2.scaffold_4690 9350782 94 - 18748788 AUGUGUGAGGGUGAGUGUAAGUGUGUUUGGGUAUG-----CAAUGCUUAGCAUAAACACCUGAACAAAGGUGUGCGGC-GUCUCCGUGUGCGCGUGUGGA---------- ((((((((.((.((...(((((((((........)-----).)))))))((....((((((......))))))...))-.)).)).).))))))).....---------- ( -25.90, z-score = 0.41, R) >dp4.chrXL_group1e 8283382 90 - 12523060 ------------AUGUAUGUAUG-G--UAUAUAUAUAUAGUAAUGCAUGU-----GUGCUCAUGUGAGUGUGUGCACCUAUCGUCAGGUGUGCAUGUGUUUGUGUUCUGA ------------.(((((((((.-.--.)))))))))(((.((((((.(.-----.(((((....))))(((..(((((......)))))..))))..).))))))))). ( -25.90, z-score = -0.81, R) >droPer1.super_13 2010090 92 + 2293547 ------------AUGUAUGUAUG-GUAUAUAUAUAUAUAGUAAUGCAUGU-----GUGCUCAUGUGAGUGUGUGCACCUAUCCUCAGGUGUGCAUGUGUUUGUGUUCUGA ------------.((((((((((-...))))))))))(((.((((((.(.-----.(((((....))))(((..(((((......)))))..))))..).))))))))). ( -26.90, z-score = -1.10, R) >consensus ______GCGUGAGUGUGUAAGUGGGUAUGGGUAUGCUUAACAAUGCCUA______ACACCUGAACAAAGGUGUGCGGC_AUCUCCGUGUGCGCGUGCAAA__________ ..............(((((..((....(((((((........)))))))......((((((......))))))...........))..)))))................. (-13.73 = -12.41 + -1.32)

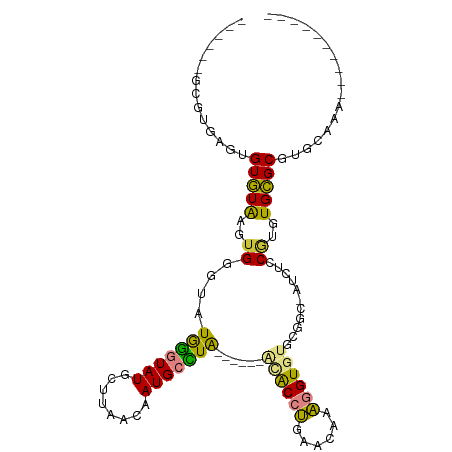
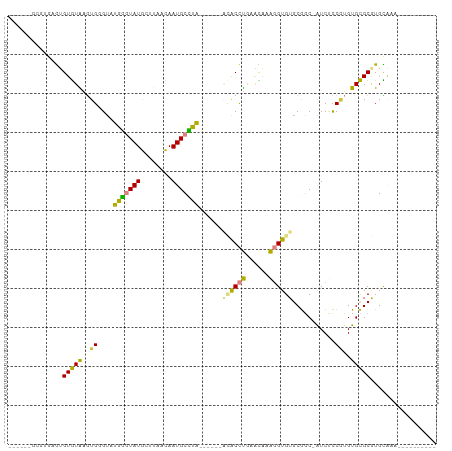
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:59:28 2011