
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,954,111 – 11,954,213 |
| Length | 102 |
| Max. P | 0.949915 |

| Location | 11,954,111 – 11,954,213 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 76.19 |
| Shannon entropy | 0.49088 |
| G+C content | 0.34510 |
| Mean single sequence MFE | -14.86 |
| Consensus MFE | -11.02 |
| Energy contribution | -10.88 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.949915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
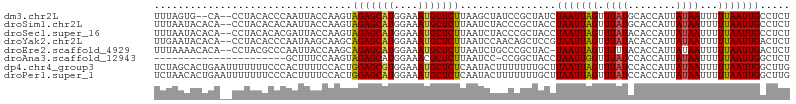
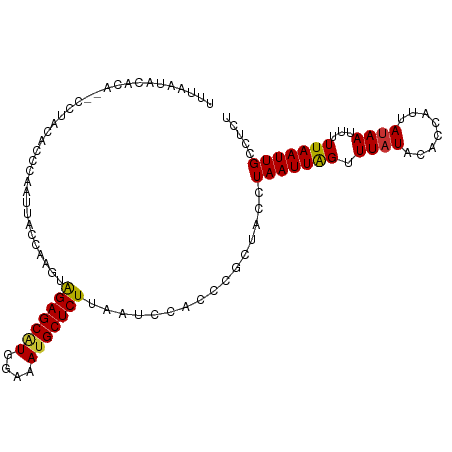
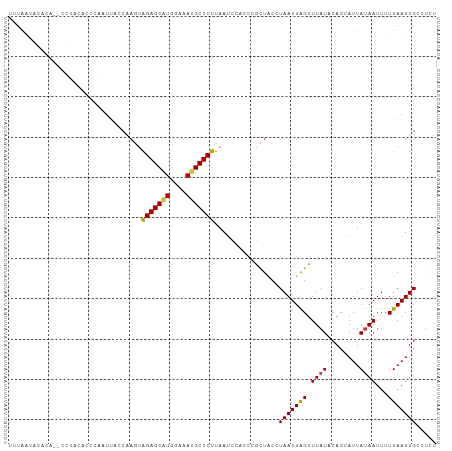
>dm3.chr2L 11954111 102 - 23011544 UUUAGUG--CA--CCUACACCCAAUUACCAAGUAGAGCAUGGAAAUGCUCUUAAGCUAUCCGCUAUCUAAUUAGUUUAUGCACCAUUAUAAUUUUUAAUUGCCUCU ....(((--((--....................(((((((....)))))))(((((((.............))))))))))))....................... ( -20.12, z-score = -2.20, R) >droSim1.chr2L 11761153 104 - 22036055 UUUAAUACACA--CCUACACACAAUUACCAAGUAGAGCAUGGAAAUGCUCUUAAUCUACCCGCUACCUAAUUAGUUUAUGCACCAUUAUAAUUUUUAAUUGCCUCU ...........--........((((((...((.(((((((....)))))))....))....((((......))))....................))))))..... ( -13.90, z-score = -1.68, R) >droSec1.super_16 153384 104 - 1878335 UUUAAUACACA--CCUACACACGAUUACCAAGUAGAGCAUGGAAAUGCUCUUAAUCUACCCGCUACCUAAUUAGUUUAUACACCAUUAUAAUUUUUAAUUGCCUCU ...........--.........(((((......(((((((....))))))))))))...........(((((((.(((((......)))))...)))))))..... ( -14.90, z-score = -2.61, R) >droYak2.chr2L 8375512 104 - 22324452 UUGAAUACACA--CCUACACCCAAUAAGCAAGCAGAGCAUGGAAAUGCUCUUAAUCCAACAGCUCCGUAAUUAGUUUAUACACCAUUAUAAUUUUUAAUUGACUCU ...........--.................((((((((((....)))))))..........)))...(((((((.(((((......)))))...)))))))..... ( -13.50, z-score = -0.91, R) >droEre2.scaffold_4929 13185286 103 + 26641161 UUUAAAACACA--CCUACGCCCAAUUACCAAGCAGAGCAUGGAAAUGCUCUUAAUCUGCCCGCUAC-UAAUUAGUUUUUACACCAUUAUAAUUUUUAAUUGACUCU ...........--........((((((......(((((((....)))))))..........((((.-....))))....................))))))..... ( -14.10, z-score = -1.68, R) >droAna3.scaffold_12943 2750812 83 + 5039921 ----------------------GCUUUCCAAGUAGAGCAUGGAAACGCUCUUAAUCC-CCGGCUACCUAAUUGGUUUAUCCACCAUUAUAAUUUUUAAUUGGCUCU ----------------------.....((((..(((((..(....))))))......-..((..(((.....)))....)).................)))).... ( -13.80, z-score = -0.13, R) >dp4.chr4_group3 4369928 106 + 11692001 UCUAGCACUGAAUUUUUUUCCCACUUUUCCACUGGAGCGUGGAAAUGCUCUCAAUACUUUUUUUGCUUAAUUAGUUUAUCCACCAUUAUAAUUUUUAAUUGGCUUG ...((((..(((.....))).....(((((((......)))))))))))...............(((.((((((.((((........))))...)))))))))... ( -14.50, z-score = -0.39, R) >droPer1.super_1 1515302 106 + 10282868 UCUAACACUGAAUUUUUUUCCCACUUUUCCACUGGAGCAUGGAAAUGCUCUCAAUACUUUUUUUGCUUAAUUAGUUUAUCCACCAUUAUAAUUUUUAAUUGGCUUG ................................((((((((....)))))).))...........(((.((((((.((((........))))...)))))))))... ( -14.10, z-score = -0.96, R) >consensus UUUAAUACACA__CCUACACCCAAUUACCAAGUAGAGCAUGGAAAUGCUCUUAAUCCACCCGCUACCUAAUUAGUUUAUACACCAUUAUAAUUUUUAAUUGCCUCU .................................(((((((....)))))))................(((((((.((((........))))...)))))))..... (-11.02 = -10.88 + -0.14)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:33:59 2011