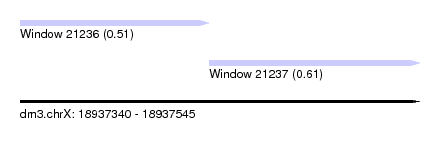
| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,937,340 – 18,937,545 |
| Length | 205 |
| Max. P | 0.614979 |

| Location | 18,937,340 – 18,937,437 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 70.78 |
| Shannon entropy | 0.38623 |
| G+C content | 0.43712 |
| Mean single sequence MFE | -19.67 |
| Consensus MFE | -16.51 |
| Energy contribution | -16.07 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -0.65 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.507377 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
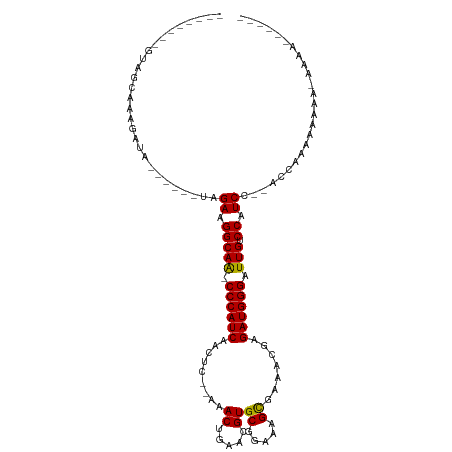
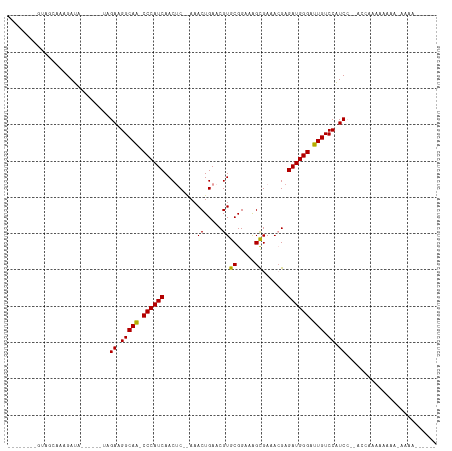
>dm3.chrX 18937340 97 + 22422827 --------GUAGCAAAGAUA------UAGAAGGCAA-CCCAUCAACUCUGAAACUGAACGUGCGGAAAGUGAAACGAGAUGGGAUUGUCCAUCCCAUCCACAAAAAAAAAAA------ --------........(((.------..((.(((((-((((((...((.......)).(((((.....))...))).)))))).))).)).))..)))..............------ ( -16.60, z-score = -0.38, R) >droYak2.chrX 17543718 117 + 21770863 GCGGCAAAGAUACAAGGAUACUCGCUUAGAAGGCAAACCCAUCAACGC-AAAACUGAACGUGCGGAAGGCGAAAGGAGAUGGGAUUGUCCAUCGUUCACCAAAAAUACAAAAUACAAA ..((....(((....(((((...(((.....)))...((((((..(((-(..........)))).....(....)..))))))..)))))))).....)).................. ( -24.80, z-score = -0.92, R) >droSim1.chrX_random 4903038 88 + 5698898 --------GUAGCAAAGAUA------UAGAAGGCAG-CCCAUCAACUC--AAACUGAACGUGCGGAAAGCGAAACGAGAUGGGAUUGUCCGUCC--ACCAAAAAAAA----------- --------............------..((.(((((-((((((...((--.....)).(((((.....))...))).)))))).))).)).)).--...........----------- ( -17.60, z-score = -0.64, R) >consensus ________GUAGCAAAGAUA______UAGAAGGCAA_CCCAUCAACUC__AAACUGAACGUGCGGAAAGCGAAACGAGAUGGGAUUGUCCAUCC__ACCAAAAAAAA_AAAA______ ............................((.(((((.((((((.........((.....))((.....)).......)))))).))).)).))......................... (-16.51 = -16.07 + -0.44)

| Location | 18,937,437 – 18,937,545 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 63.36 |
| Shannon entropy | 0.58297 |
| G+C content | 0.41650 |
| Mean single sequence MFE | -26.44 |
| Consensus MFE | -9.05 |
| Energy contribution | -9.43 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.614979 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
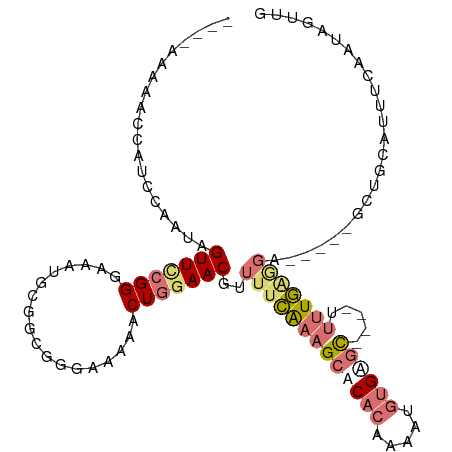
>dm3.chrX 18937437 108 + 22422827 AAACAAAAACCAUCCAAUAGUUCCGGGAAAUGCGGUGGAAAAAACUGGAACGUUUUUAAAGCACACAAAAUGUGAGCCGCGUUUUAAAGA-----GCCAUGUUUCAAUAGUUG ...(((..........((.((((...((((((((((...((((((......))))))....((((.....)))).))))))))))...))-----)).))..........))) ( -22.85, z-score = -0.56, R) >droEre2.scaffold_4690 9220160 96 + 18748788 AAAAAGAAACCAUCCGAUAGUUCCGGGAAAUGC--UGGGAAAAACUGGAACAUUUUCGUAGCUC-CGUUCUAGGGAU----UCCUGGGGCUCUU-GCUCAGCUU--------- ..............(((.(((((((((((..((--((.(((((.........))))).))))((-(......))).)----)))))))))).))-)........--------- ( -27.40, z-score = -0.59, R) >droSec1.super_8 1231210 93 + 3762037 ----------------AUAGUUCGGGGAAAUGUGGCGGGAAAAACUGAAACGUUUUCAAAGCUCACAAAAUGUGAGC----UUUUGAGGAUUUAGGCUGCAUUUCAAUAGUUG ----------------..........((((((..((((......)).....(((((((((((((((.....))))))----.)))))))))....))..))))))........ ( -30.30, z-score = -4.22, R) >droSim1.chrX_random 4903126 95 + 5698898 ----GAAAACCAUCCAAUAGUUUCGGGAAAUGCGGCGGGAAAAACUGGAACGUUUUCAAAGCACACAAAAUGUGAGC----UUUUG----------CUGCAUUUCAAUAGUUG ----......................(((((((((((.(((((.(......))))))(((((.(((.....))).))----)))))----------)))))))))........ ( -25.20, z-score = -1.87, R) >consensus ____AAAAACCAUCCAAUAGUUCCGGGAAAUGCGGCGGGAAAAACUGGAACGUUUUCAAAGCACACAAAAUGUGAGC____UUUUGAGGA_____GCUGCAUUUCAAUAGUUG .........................(((((((...(((......)))...)))))))...((((((.....)))))).................................... ( -9.05 = -9.43 + 0.37)

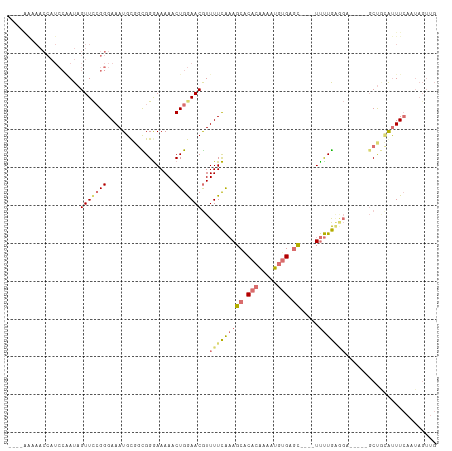
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:58:50 2011