| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,868,911 – 18,869,060 |
| Length | 149 |
| Max. P | 0.946862 |

| Location | 18,868,911 – 18,869,060 |
|---|---|
| Length | 149 |
| Sequences | 7 |
| Columns | 170 |
| Reading direction | forward |
| Mean pairwise identity | 80.11 |
| Shannon entropy | 0.37649 |
| G+C content | 0.46674 |
| Mean single sequence MFE | -32.76 |
| Consensus MFE | -24.14 |
| Energy contribution | -26.06 |
| Covariance contribution | 1.92 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.946862 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
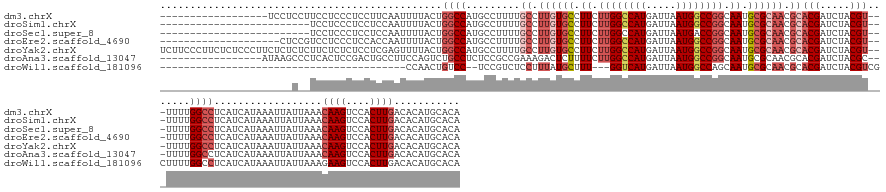
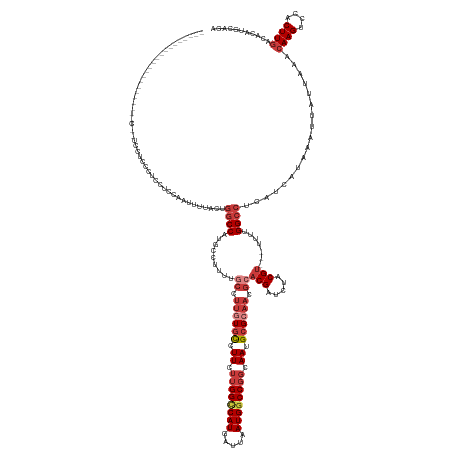
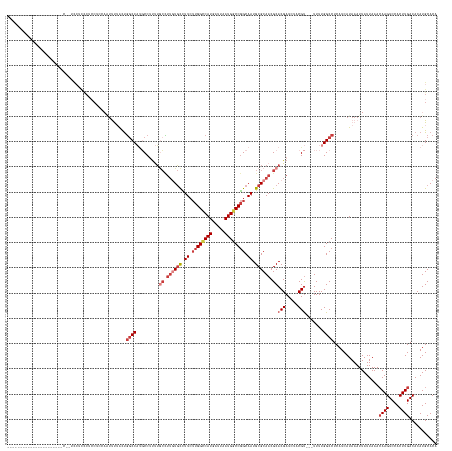
>dm3.chrX 18868911 149 + 22422827 ------------------UCCUCCUUCCUCCCUCCUUCAAUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGU---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA ------------------.............................(((((........((.((((((.((.((((((((.....)))))))).)).)))))).))(((.....)))---...)))))..................((((....))))........... ( -35.30, z-score = -2.62, R) >droSim1.chrX 14579092 142 + 17042790 -------------------------UCCUCCCUCCUCCAAUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGU---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA -------------------------......................(((((........((.((((((.((.((((((((.....)))))))).)).)))))).))(((.....)))---...)))))..................((((....))))........... ( -35.30, z-score = -2.51, R) >droSec1.super_8 1164078 142 + 3762037 -------------------------UCCUCCCUCCUCCAAUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGACCGGCAAUGCGCAACGCACGAUCUACGU---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA -------------------------......................(((((........((.((((((.((.((((.(((.....))).)))).)).)))))).))(((.....)))---...)))))..................((((....))))........... ( -28.20, z-score = -0.97, R) >droEre2.scaffold_4690 9151488 147 + 18748788 --------------------CUCCGUCCUCCCUCCACCAAUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGU---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA --------------------...........................(((((........((.((((((.((.((((((((.....)))))))).)).)))))).))(((.....)))---...)))))..................((((....))))........... ( -35.30, z-score = -1.97, R) >droYak2.chrX 17477242 167 + 21770863 UCUUCCCUUCUCUCCCUUCUCUCUCUUCUCUCUCCUCGAGUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGU---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA .............................(((.....))).......(((((........((.((((((.((.((((((((.....)))))))).)).)))))).))(((.....)))---...)))))..................((((....))))........... ( -36.30, z-score = -2.77, R) >droAna3.scaffold_13047 1754745 150 + 1816235 -----------------AUAAGCCCUCACUCCGACUGCCUUCCAGUCUGCCUCUCCGCCGAAAGACUCUUUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGC---UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA -----------------...............(((((.....)))))(((......((((((((.......((((((((((.....))))))))...((((...)))).))......)---)))))))...................((((....))))......))).. ( -35.22, z-score = -2.08, R) >droWil1.scaffold_181096 8090735 124 + 12416693 -----------------------------------------CCAACUGUCC--UCCGUCUCCUUUAUGCUUU---GGUCAUGAUUAAUGGCCAGCAAUGCGCAACGCACGAUCUACGUCGCUUUUGGCCUCAUCAUAAAUUAUUAAAGAAGUCCACUUGACACAUGCACA -----------------------------------------..........--.............(((..(---(((((((((....(((((((.....))...(((((.....))).))...)))))..)))))............(((....)))))).)).))).. ( -23.70, z-score = 0.09, R) >consensus ______________________C__UCCUCCCUCCUCCAAUUUUACUGGCCAUGCCUUUUGCCUUGUGCCUUCUUGGCCAUGAUUAAUGGCCGGCAAUGCGCAACGCACGAUCUACGU___UUUUGGCCUCAUCAUAAAUUAUUAAACAAGUCCACUUGACACAUGCACA ...............................................((((.........((.((((((.((.((((((((.....)))))))).)).)))))).))(((.....))).......))))..................((((....))))........... (-24.14 = -26.06 + 1.92)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:58:31 2011