
| Sequence ID | dm3.chrX |
|---|---|
| Location | 18,841,533 – 18,841,644 |
| Length | 111 |
| Max. P | 0.705717 |

| Location | 18,841,533 – 18,841,644 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 78.64 |
| Shannon entropy | 0.40259 |
| G+C content | 0.47222 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -17.75 |
| Energy contribution | -19.59 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.635581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18841533 111 + 22422827 ----AGUCCCAGGCAGAACCUUUAACGUUCUAUCGACAAAAGUCUCAUUGUCUCACGGGCCACAUGGCCCUGUAAUCAAAUAUCAAUGAGAUGAAGGAGGUACUAUCAUUCUGCA ----........(((((((((((..((......))......(((((((((......(((((....)))))((....)).....)))))))))))))).(((...))).)))))). ( -32.50, z-score = -2.55, R) >droSim1.chrX_random 4891541 111 + 5698898 ----AGUCCCAGGCAGAACCUUUAACGGUCUACCGACAAAAGUCUCAUUGUCUCACGGGCCACAUGGCCCUGUAAUCAAAUGUCAAUGAGAUGAAGGAGGUACUAUCAUUCUGCA ----........(((((((((((..(((....)))......(((((((((.(....(((((....)))))((....))...).)))))))))))))).(((...))).)))))). ( -37.20, z-score = -3.09, R) >droSec1.super_8 1145691 111 + 3762037 ----AGUCCCAGGCAGAACCUUUAACGGCCUACCGACAAAAGUCUCAUUGUCUCACGGGCCACAUGGCCCUGUAAUCAAAUGUCAAUGAGAUGAAGGACGUACUAUCAUUCUGCA ----........(((((((((((..(((....)))......(((((((((.(....(((((....)))))((....))...).))))))))))))))..(......).)))))). ( -35.90, z-score = -3.09, R) >droYak2.chrX 17458456 111 + 21770863 ----AGUUUCAGGCAGAAUCUUUAACGGUCUACCGACAAAAAUCUCAUUGUCUCACGGAGCUCGUGGCCCUGCAAUCAAAUAUCAAUGAGAUGGAGGAGGCACUAUCAUCCUGCA ----........((((...........((((.((..((....((((((((...((((.....))))((...))..........))))))))))..)))))).........)))). ( -25.65, z-score = 0.23, R) >droEre2.scaffold_4690 9133453 111 + 18748788 ----AGUCCCAGGCAGAACCUUUAACGGUCUAUCGACAAAAGUCUCAUUGUCUCACGGGCCACAUGGCCCUGCAAUCAAAUAUCAAUGAGAUGGAGGCGGUACUAUCAUCCUGCA ----........((((.(((......))).(((((.(....(((((((((...((.(((((....)))))))...........)))))))))....))))))........)))). ( -32.24, z-score = -1.52, R) >droVir3.scaffold_12726 779741 100 + 2840439 CAACAGCGUGCCACGCCGUCGCCAGCGUGUCUUUGACAAAAGGCUCAUUGUUUUGC----CAUAUGGUCCCGUAAUCAAAUAUCAAUG----GAAGGGUCCACUUCCA------- .......(.(((((((........))))(((...)))....))).)((((.((((.----..((((....))))..))))...))))(----((((......))))).------- ( -23.10, z-score = 0.27, R) >consensus ____AGUCCCAGGCAGAACCUUUAACGGUCUACCGACAAAAGUCUCAUUGUCUCACGGGCCACAUGGCCCUGUAAUCAAAUAUCAAUGAGAUGAAGGAGGUACUAUCAUUCUGCA ...........(..((.((((((..(((....)))......(((((((((...((.(((((....)))))))...........)))))))))...)))))).))..)........ (-17.75 = -19.59 + 1.84)


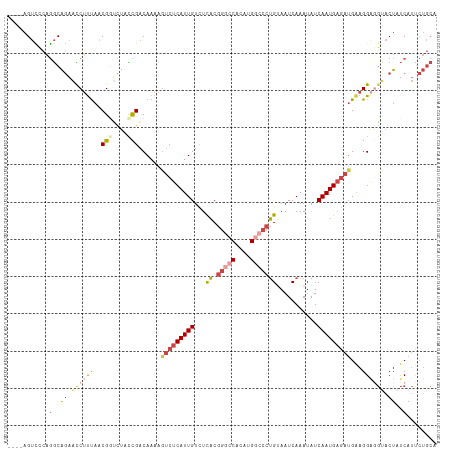
| Location | 18,841,533 – 18,841,644 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 78.64 |
| Shannon entropy | 0.40259 |
| G+C content | 0.47222 |
| Mean single sequence MFE | -33.25 |
| Consensus MFE | -20.89 |
| Energy contribution | -22.50 |
| Covariance contribution | 1.61 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.705717 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 18841533 111 - 22422827 UGCAGAAUGAUAGUACCUCCUUCAUCUCAUUGAUAUUUGAUUACAGGGCCAUGUGGCCCGUGAGACAAUGAGACUUUUGUCGAUAGAACGUUAAAGGUUCUGCCUGGGACU---- .(((((((................((((((((........((((.(((((....)))))))))..))))))))((((...((......))..)))))))))))........---- ( -32.20, z-score = -1.56, R) >droSim1.chrX_random 4891541 111 - 5698898 UGCAGAAUGAUAGUACCUCCUUCAUCUCAUUGACAUUUGAUUACAGGGCCAUGUGGCCCGUGAGACAAUGAGACUUUUGUCGGUAGACCGUUAAAGGUUCUGCCUGGGACU---- .(((((((................((((((((.(....).((((.(((((....)))))))))..))))))))((((...(((....)))..)))))))))))........---- ( -36.60, z-score = -2.17, R) >droSec1.super_8 1145691 111 - 3762037 UGCAGAAUGAUAGUACGUCCUUCAUCUCAUUGACAUUUGAUUACAGGGCCAUGUGGCCCGUGAGACAAUGAGACUUUUGUCGGUAGGCCGUUAAAGGUUCUGCCUGGGACU---- ................(((((...((((((((.(....).((((.(((((....)))))))))..))))))))........((((((((......)).)))))).))))).---- ( -38.80, z-score = -2.41, R) >droYak2.chrX 17458456 111 - 21770863 UGCAGGAUGAUAGUGCCUCCUCCAUCUCAUUGAUAUUUGAUUGCAGGGCCACGAGCUCCGUGAGACAAUGAGAUUUUUGUCGGUAGACCGUUAAAGAUUCUGCCUGAAACU---- .(((((((...............(((((((((.............((((.....))))(....).)))))))))(((((.(((....))).))))))))))))........---- ( -28.60, z-score = -0.46, R) >droEre2.scaffold_4690 9133453 111 - 18748788 UGCAGGAUGAUAGUACCGCCUCCAUCUCAUUGAUAUUUGAUUGCAGGGCCAUGUGGCCCGUGAGACAAUGAGACUUUUGUCGAUAGACCGUUAAAGGUUCUGCCUGGGACU---- .(((((((................((((((((........(..(.(((((....))))))..)..))))))))((((.(((....)))....)))))))))))........---- ( -32.50, z-score = -0.72, R) >droVir3.scaffold_12726 779741 100 - 2840439 -------UGGAAGUGGACCCUUC----CAUUGAUAUUUGAUUACGGGACCAUAUG----GCAAAACAAUGAGCCUUUUGUCAAAGACACGCUGGCGACGGCGUGGCACGCUGUUG -------((((((......))))----))......((((((...(((..(((.((----......)))))..)))...))))))..(((((((....)))))))........... ( -30.80, z-score = -1.58, R) >consensus UGCAGAAUGAUAGUACCUCCUUCAUCUCAUUGAUAUUUGAUUACAGGGCCAUGUGGCCCGUGAGACAAUGAGACUUUUGUCGAUAGACCGUUAAAGGUUCUGCCUGGGACU____ .(((((((................((((((((.....((....))(((((....)))))......))))))))((((...((......))..)))))))))))............ (-20.89 = -22.50 + 1.61)


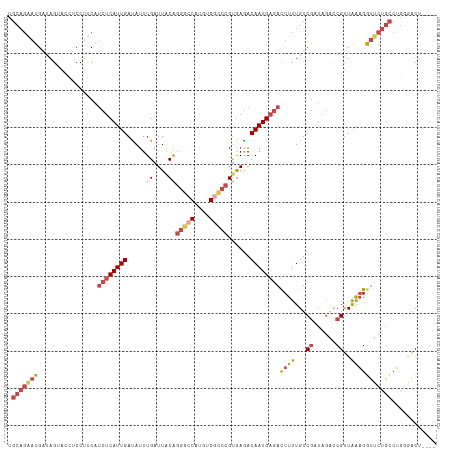
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:58:26 2011